
rmoretti Staff Lv 1
We’re pleased to announce that the CACHE organizers have officially released the results from the CACHE Challenge #2, and Foldit has done notably well. (Press Release, technical details)
*We're hoping to publish a paper about the CACHE challenge #2 – If you participated in the puzzles and wish to be listed as part of the Foldit Players consortium, please fill out this form.*
CACHE Challenge #2 was the first CACHE challenge Foldit participated in and was the one targeting the NSP13 helicase domain of SARS-CoV-2 (blog link). In particular, the Challenge proceeded in two phases, the first being generation of a set of possible compounds predicted from a Compound Library, and the second being a “refinement” step, where participants were asked to take hit compounds from which showed activity in the first phase and come up with related compounds which also showed (hopefully better) activity.
Phase 1 Approach
For Phase 1, Foldit ran a series of eight puzzles. These puzzles were the first to have the Compound Library tool enabled, allowing players to search for compounds within the Enamine REAL compound library, from which the CACHE organizer would order compounds. In the eight puzzles, over 150 players submitted at least one compound to the server, resulting in a total of 7598 potential compounds, 1411 of which were part of the compound library. (The Compound Library Bonus was only applied for rounds 7 & 8). For each compound, we took protein ligand complex with the highest Foldit score as representative.
To select the compounds to submit, we then took all the library compounds submitted and redocked them using the RosettaLigand docking program. The goal here was to find compounds which looked to be stable in the submitted conformation and weren’t precariously perched in an artifact of the scoring function. We originally hoped to use a machine-learning based approach to provide an orthogonal assessment of compound binding, but we saw that this approach didn’t work well with the Foldit-produced structures. (Better assessment of structure results is something we’re still working on.) Compounds were evaluated both on their predicted binding energy as well as how closely the stayed to the player-designed structure. Approximately 150 structures were initially selected, and after checking with Enamine for cost and availability, this was reduced to a final submission of 111 compounds.
Phase 1 Results
Of the 111 Foldit-derived compounds ordered, 76 were successfully synthesized. These compounds were tested by CACHE organizers and their collaborators. Specifically, they used a technique called Surface Plasmon Resonance (SPR) to look at how well each compound bound to the protein of interest. They also used Dynamic Light Scattering (DLS) to figure out if the compounds were well behaved (i.e. were soluble and didn’t aggregate).
Two Foldit derived compounds (out of 46 total from 18 groups) were deemed by the CACHE organizers to be worth taking through to the next round (reported here). Once again, congratulations to Aubade01 and an anonymous player for their designs.
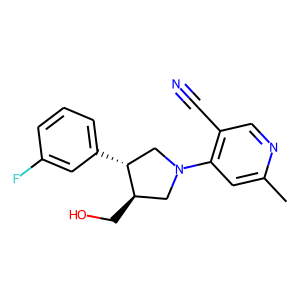
Hit 1; CACHE_1414_40
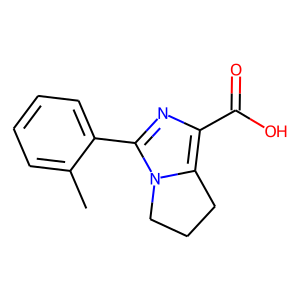
Hit 2; CACHE_1414_34
Phase 2 Approach
Following up on the positive results from phase 1, we ran a series of four puzzles which asked players to diversify the two hit compounds (link). Puzzle setup was similar to before, but with the addition of the compound similarity. The goal in this round is not necessarily to come up with better binding compounds (but that’s hoped for), but instead to help establish what’s called “structure-activity relationship”. The concept is that true binders aren’t entirely unique. Instead, they sit in “well” in compound space, where small changes to the compound structure don’t have too much of an affect on binding. If it’s a true binder (as opposed to an artifact of the assay), then there should be several closely related compounds which also should be binders.
Selection of player designs proceeded similarly to phase 1 – the only difference was that in addition to the in-library filter, there was also a compound similarity filter – though the in-game objective meant that basically all good-scoring compounds passed this filter. From the four rounds, 2413 total compounds were obtained, 948 of which were in the compound library. This time, 75 structures were initially selected, which was reduced to 48 compounds due to cost and availability.
Phase 2 Results
Of the 48 compounds ordered, 34 were successfully synthesized and tested. This time, in addition to the SPR and DLS testing, the CACHE organizers double checked that the compounds were binding to the desired site with Nuclear Magnetic Resonance (NMR) measurements, at least for a subset of compounds which contained fluorine atoms. (Due to the particular experiment they were using, they can only “see” fluorine with the NMR.)
Of the 34 compounds, 10 were analogs of Hit 2/CACHE_1414_34. However, none provided the type of binding the CACHE organizers were looking for. The 24 remaining compounds were analogs of Hit 1/CACHE_1414_40. Three of these compounds showed the desired binding results.
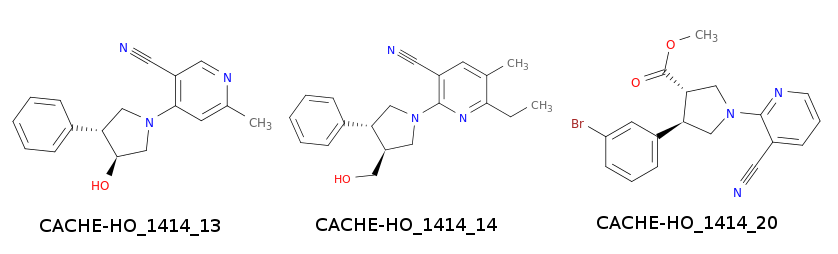
All three compounds came from Round 3. Congratulations to blazegeek, Bletchley Park, & AlphaFold2 for coming up with these compounds.
How Foldit did
It’s instructive to take a look at how well the Foldit design process works in comparison to other participants. There’s a number of graphs at the CACHE website which compare the performance of the various groups (Foldit is participant 1414) – Foldit players have the highest scoring single molecule (Hit 1/CACHE_1414_40), and are ranked in fourth place (out of 22) when the total number of active molecules are considered.
Most notably, CACHE_1414_40 and the active derivatives (along with 6 other compound groups by other participants) is considered by the CACHE organizers to be “promising [and] should be progressed into more potent molecules to investigate anti-viral activity.” This is particularly exciting, as this target (the helicase) shows promise as a pan-coronavirus target, meaning drugs developed from these molecules could potentially be active across all coronaviruses, not just SARS-CoV-2.
Phase 1.5 Approach & Results
I should also mention that we were also asked to rank all the compounds that were submitted by every participant. For CACHE Challenge #2, we unfortunately did not have things set up to have players do that ranking. Instead, we effectively did the post ranking selection process, but scoring purely based on RosettaLigand docking energy. On this (automated) basis, the group came in 3rd place for accuracy of predicting compound binding strength. (Figure 3 at the CACHE results website.
CACHE Challenge #3
Foldit has also participated in CACHE Challenge #3, and the CACHE organizers are still processing the results. While we can’t give a full update until the results are officially released hopefully later this fall, preliminary indications are that Foldit also did rather well in that challenge - stay tuned for updates.