
rmoretti Staff Lv 1
You may remember that approximately a year ago we ran a series of puzzles which looked to find small molecules which bound to the KLHDC2 E3 ligase. Like VHL, (for which we've had success), molecules which bind KLHDC2 would be helpful in creating PROTACs, a new class of molecular regulators. But unlike VHL, there isn't a good druglike molecule which binds KLHDC2, just the naturally occurring peptide.
Collaborators at Boehringer Ingelheim have pored over the results of the first set of KLHDC2 puzzles. Due to logistics considerations, custom synthesis of the full designs wasn't feasible at the time. Additionally, the direct path from peptide to small molecule is a difficult and risky one. Instead, based on the first round results, medicinal chemists identified 17 core motifs which were common in the designed molecules, looked promising for further development, and which could be easily obtained. By confirming which compound cores which work well, we can have better confidence that derivative molecules would show activity.
BI experimentalists then used a technique called saturation transfer difference NMR, which is sensitive to even weak binding. They could confirm binding of a number of molecules, and could even confirm that they bound in the proper location by examining how the naturally occurring peptide competed with small molecule binding.
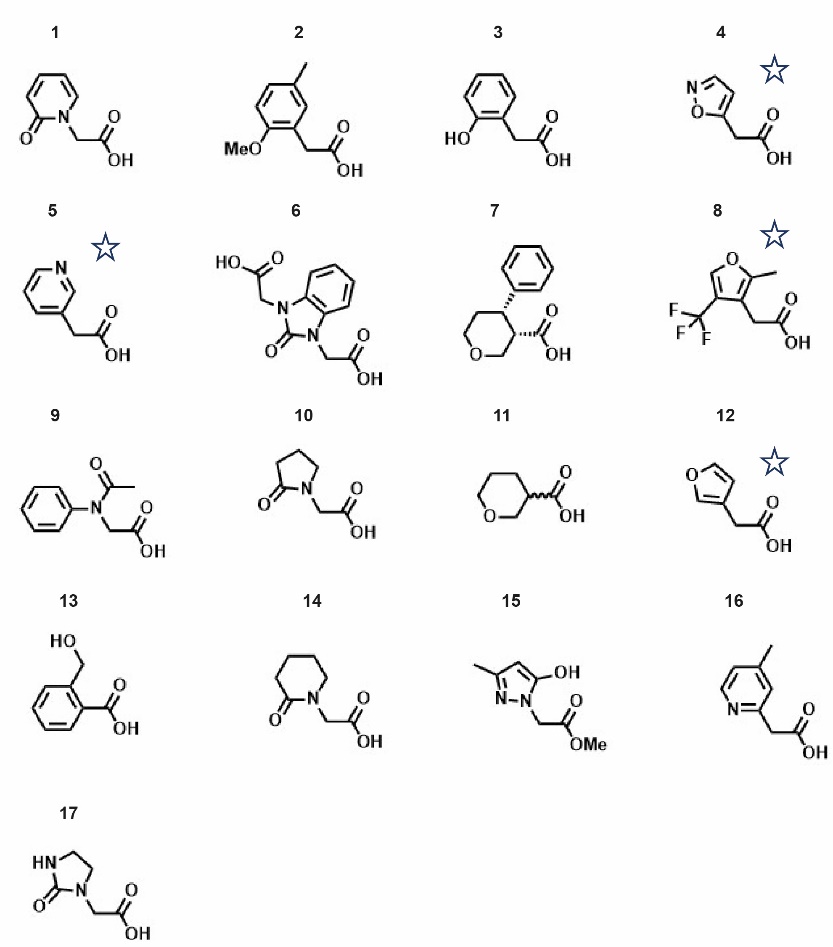
A listing of all the fragment molecule tested. The compounds with stars indicate those which showed binding activity.
With the knowledge of which compounds bound, we embarked on the sometimes difficult process of determining crystal structures of the bound complex. And we were successful! Crystals with compound 12 bound to the KLHDC2 protein were obtained, and they showed that binding happened in exactly the way we hoped.
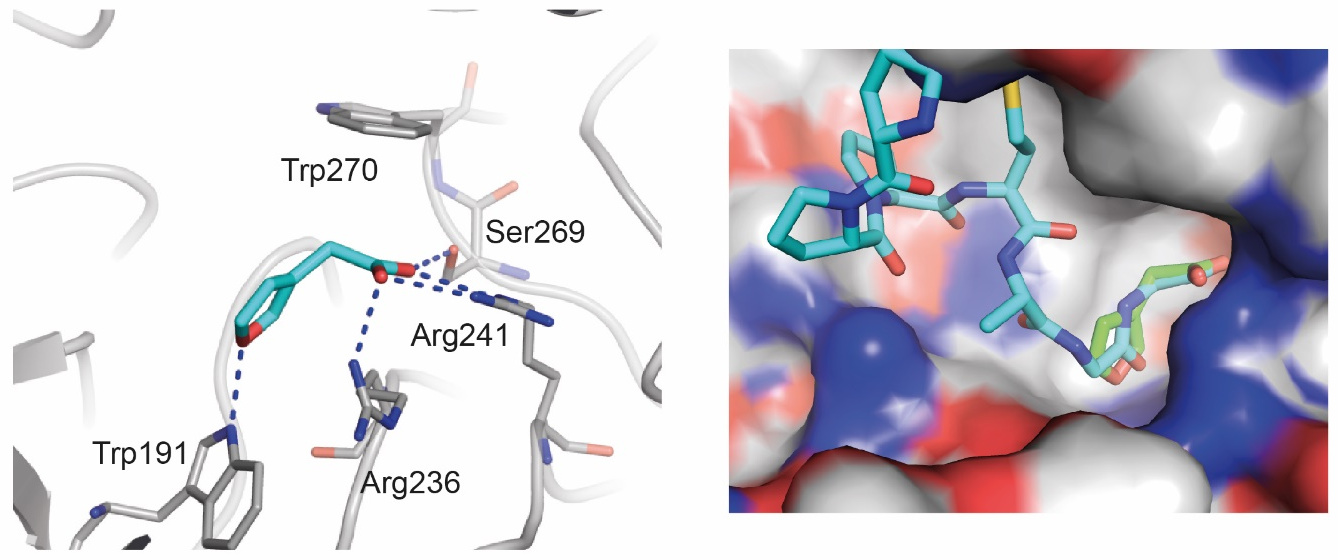
Binding of the small molecule to the KLHDC2 protein. Left, the major interactions of the compound to the protein. Right, a comparison of the small molecule (in green) with the binding mode of the naturally occurring peptide (cyan).
Enter KLHDC2: the Next Level
Based on these exciting results, we're launching a new series of KLHDC2 puzzles. This time around, we want to start with the discovered small molecule. Keep the core structure and the existing hydrogen bonds, but build up the structure, making additional interactions with the protein.




