
rmoretti Staff Lv 1
The Critical Assessment of Computational Hit-Finding Experiments (CACHE) challenge has recently announced their next challenge target, and Foldit players have once again been selected to participate in finding small molecule inhibitors.
Foldit has previously participated in CACHE challenge #2 (blog post). The identity of the 100 or so molecules discovered by Foldit players were submitted to the CACHE organizers at the end of November, and those molecules (along with several thousand others from other CACHE participant groups) are currently being ordered and tested by CACHE's experimental collaborators. We anticipate having the results from those experimental tests sometime this summer.
But while those compounds are being tested, we have a new target for which to predict small molecule binders:
CACHE Challenge #3 – SARS-CoV-2 Nsp3 macrodomain
Like the previous challenge, next CACHE challenge focuses on finding inhibitors for SARS-CoV-2 enzymes. This time it's an enzyme which doesn't normally get much attention – the "macrodomain" from "nonstructural protein 3 (Nsp3)". Nsp3 is a large protein with multiple domains, and one of them has ADP-ribosyl hydrolase activity.
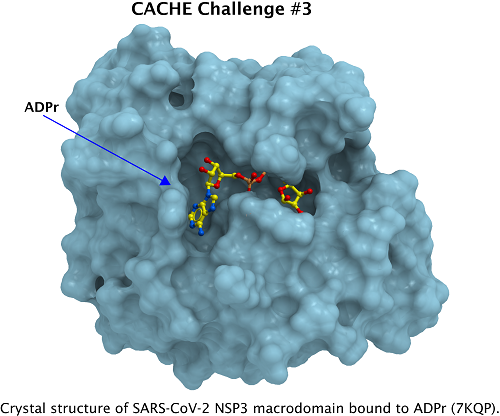
Image courtesy of cache-challenge.org
The immune response to SARS-CoV-2 relies in part on ADP-ribosylation of certain proteins - that is, certain enzymes in the immune system decorate proteins with ADP-ribose groups. The Nsp3 macrodomain from the virus removes these groups. In effect, the ADP-ribose groups function as "Warning: Virus!" posters which your body's security team post in response to infection, and the Nsp3 macrodomain is the member of the SARS gang which runs around tearing them down. There are indications that inhibiting the Nsp3 macrodomain has the potential to greatly improve the immune systems ability to fight off a COVID infection.
The structure of the Nsp3 macrodomain in complex with its ADP-ribose substrate is known, and the protein has a well-defined substrate binding site. There's already been a bunch of work looking into various compounds which might function as inhibitors by binding in the same location as the ADP-ribose. The Protein Databank has several hundred crystal structures with various small molecules bound to the Nsp3 macrodomain. (These are available as group deposition IDs G_1002236, G_1002238 and G_1002239. (Simply put those numbers in to the search box on https://www.rcsb.org/ to find the list of structures.)
One limitation of many of these compounds is that they contain a carboxylic acid or carboxylate group. This is somewhat unsurprising, as the ADP-ribose substrate has acidic phosphate groups, and a carboxylate can mimic many of the same interactions the phosphate can make with the protein. However, having a carboxylate on the inhibitors can be a liability, as it makes it harder for the drugs to cross the cell membranes and get into cells where they need to be to function. So the CACHE organizers are particularly interested in new compound ideas which do not contain carboxylates.
Foldit and CACHE Challenge #3
Over the next month or so we'll be running a series of puzzles of small molecule design for inhibitors of Nsp3 macrodomains. As with CACHE Challenge #2, we're asking you to use the Compound Library panel to find compounds which are readily purchasable. From the results which come back from players, we'll select ~100 compounds to send to the CACHE organizers, who will then test how well they bind to and inhibit the SARS-CoV-2 Nsp3 macrodomain.
Due to limitations in synthesis, CACHE organizers will only be able to obtain and test library compounds. As before, we imagine an iterative approach will yield the most success. Use the small molecule design tools to create novel molecules which you think will work well. But then use the Compound Library panel to find "on-library" compounds which are similar. Use the other tools in Foldit (pull, wiggle, shake, ligand tweak) to optimize the placement of the library compound. Then go back to the small molecule design tool to discover novel modifications. Run this new molecule through library search to find close library compounds, etc. Only compounds which are available in the library will be selected. See the CACHE Challenge #2 blog post for more info about the Compound Library panel.
The compound library objective should give a bonus for compounds that are available in the library. It doesn't know about all 20+ billion compounds in the library, though, just the ones which you've downloaded from the compound library search. If you're loading a share on a different client, you may need to re-submit the compound to the library search.
Administrative
Participation in CACHE puzzles is subject to the CACHE Terms of Participation, in particular “the Challenge IP [including Challenge Compounds] will be made freely available in the public domain pursuant to Creative Commons Attribution Only (CC-BY 4.0 or subsequent versions) licensing terms, with the intent that such Challenge IP may be Used and practiced by Users for any purpose”.
Again, special thanks to John Irwin and the ZINC database for the ultra-large library compound search behind the Compound Library panel.

