
rmoretti Staff Lv 1
We have more preliminary results to share from the CACHE challenge!
The CACHE Challenge
Earlier this year we launched a puzzle series as part of the CACHE Challenge. CACHE is like CASP for small molecule drug design – it’s an independent, blind prediction task to see if computationalists can do a good job of predicting which small molecule can bind to a given protein structure. We've previously worked on CACHE Challenge #2 (SARS Helicase), and we're still awaiting results on the followup of that effort. This next Foldit CACHE puzzle series is for CACHE Challenge #3.
Since the initial puzzle series ended, we've taken all the designs which Foldit players created in the five rounds, filtered by various quality metrics as well as to make sure they were part of the compound library. Approximately 2000 compounds made the cut, and were redocked into the protein to make sure that the designed binding mode was specific. From the redocking results, we ranked the compounds, selected ~100 of them to send to the CACHE organizers for ordering and testing.
These ~100 compounds were then combined with the rest of the molecules submitted by the other ~22 participants, and those ~1700 compounds were the basis of the recent CACHE SARS Nsp3 Reranking challenge puzzle series (the results of that effort are still to be determined).
Results
Since then CACHE has ordered and tested the compounds. Due to the difficulties of chemical synthesis, not all the compounds which we submitted were able to be tested, but 81 of them were. The CACHE organizers and their collaborators did a number of different types of assays to make sure that the compounds were binding to the protein, weren’t binding nonspecifically, and didn’t have odd aggregation or other such interference.
And Foldit players designed two compounds which passed this initial screen! Congratulations to Sandrix72 and ucad for their designs. (If you'd like to be mentioned in future Foldit blog posts and papers, you can go to https://fold.it/profile/edit or click the gear icon in the upper right when logged in to change the “Foldit can share my username” setting.) Both successful compounds were from Round 2, though they weren’t the top scoring compounds from that round.
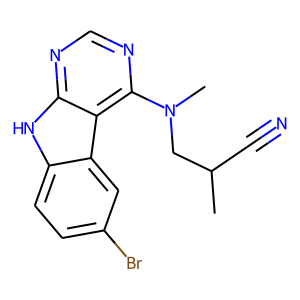
Hit 1; SMILES CC(CN(C)c1c2c3cc(ccc3[nH]c2ncn1)[Br])C#N
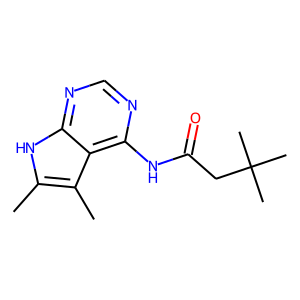
Hit 2; SMILES Cc1c2c(NC(CC(C)(C)C)=O)ncnc2[nH]c1C
Foldit did relatively well in its submission - only 6 out of 23 groups had compounds selected for being advanced to the next phase, and only 12 compounds in total were advanced. In the intial screens, both compounds have an affinity of around 40 µM, which is reasonable but not fantastic. It's possible one of the derivatives found will be better!
Onto the next phase
Since we have compounds which passed preliminary screening, we’ve been invited to participate in the next phase! In this phase, we’re asked to explore the “structure activity relationship” of the compounds we had success with. That is, can we find compounds which are similar to the compounds we submitted, but which have better binding affinity?
Similar to the first phase, only compounds which are in the compound library will be considered. Additionally, we need to submit compounds which are “close enough” to our hit compounds. There isn’t a hard threshold on this, but the intent is to make the hit compounds better, rather than come up with completely novel compounds. Also keep in mind that we don’t have experimental structures of the protein-ligand complex, so the starting location of the compound may not be where or how it actually binds.
UPDATE – Additional compounds!
The CACHE organizers have gotten back to us with good news. It turns out that Foldit players designed four additional compounds with potential activity! These compounds weren't detectable in the initial round of screening because they likely had solubility/aggregation issues under the original assay conditions. When the experimental conditions were adjusted, these additional compounds were discovered. (I should note that the Foldit group is unlikely to be alone in getting additional compounds. While the CACHE organizers haven't mentioned details, other groups have likely also picked up additional compounds.)
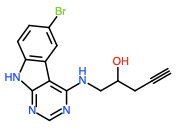
Hit 3; SMILES OC(CNC=1N=CN=C2NC=3C=CC(Br)=CC3C12)CC#C Round 2
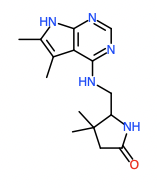
Hit 4; SMILES CC=1NC=2N=CN=C(NCC3NC(=O)CC3(C)C)C2C1C Round 3
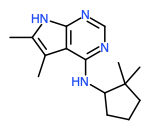
Hit 5; SMILES CC=1NC=2N=CN=C(NC3CCCC3(C)C)C2C1C Round 3
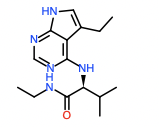
Hit 6; SMILES CCNC(=O)[C@@H](NC=1N=CN=C2NC=C(CC)C12)C(C)C Round 2
Congratulations to nspc and Bruno Kestemont for coming up with these compounds!
Since we have more compounds, we're running a few extra weeks of CACHE #3 puzzles for some of these new puzzles. We hope that these new starting points can let you explore new areas of structure/activity space, finding additional new potential compounds.
Participation in CACHE puzzles is subject to the CACHE Terms of Participation, in particular “the Challenge IP [including Challenge Compounds] will be made freely available in the public domain pursuant to Creative Commons Attribution Only (CC-BY 4.0 or subsequent versions) licensing terms, with the intent that such Challenge IP may be Used and practiced by Users for any purpose”.
