LociOiling Lv 1
I've been looking back at the ED recon puzzles, trying to identify the ones that had problems which might affect the value of the results.
A number of the ED recons had puckers or straws, where Foldit thinks there is a peptide bond between distant residues. Straws are easy to spot just by looking at the puzzle, but puckers may require looking into the details, like the ideality subscores. Puckers tend to happen where there are missing residues.
The starting pose of puzzle 2323 has an obvious straw at 65-66. The corresponding residues in 3HRY are A 65 and A 66. So there are no missing residues, but A 65 and A 66 shouldn't be bonded.
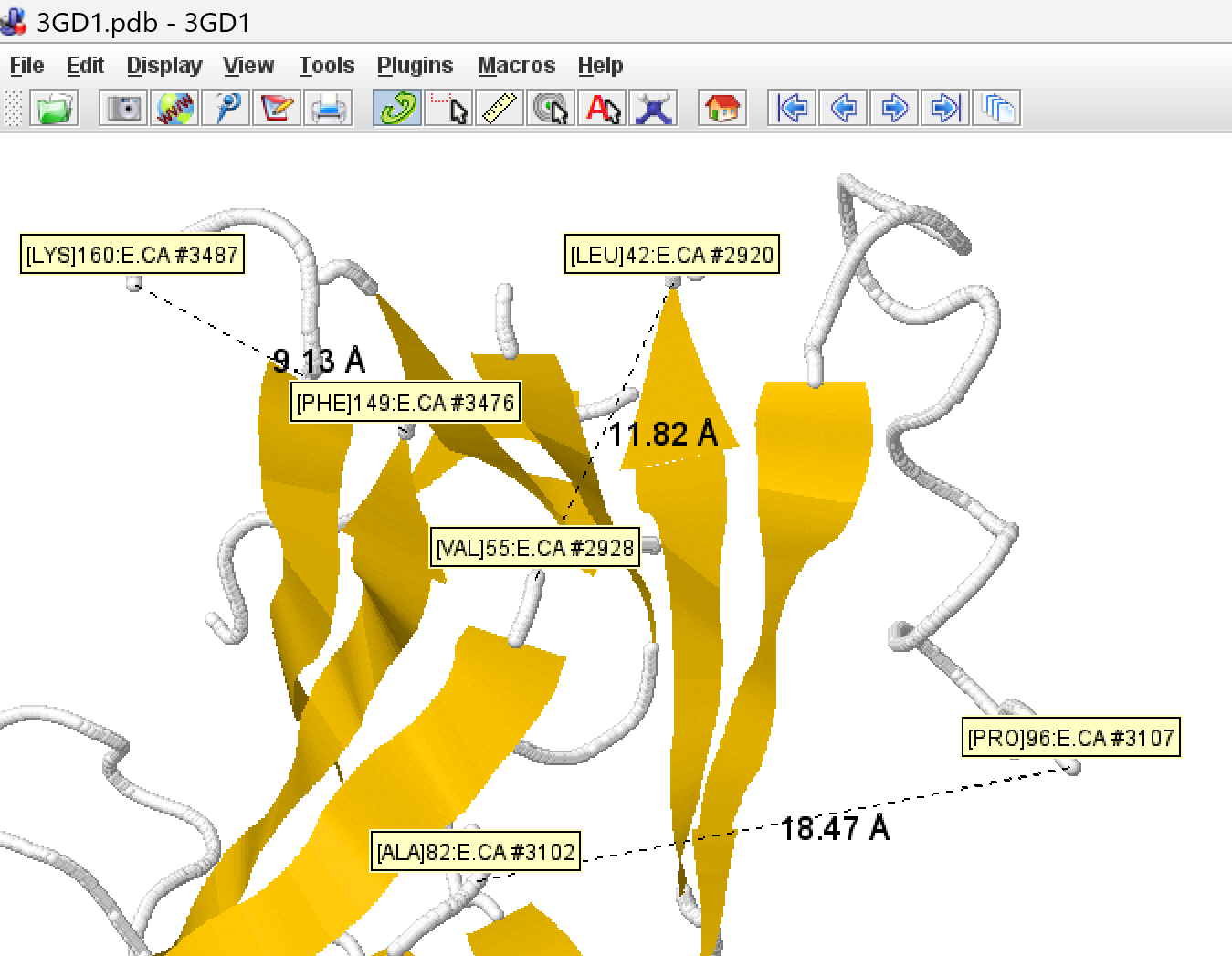
On the PDB site, the Mol* viewer shows a dotted line between A 65 and A 66. Jmol simply shows a large gap between the two, but they're clearly not connected.
Foldit thinks that 65 and 66 are connected, as shown here:
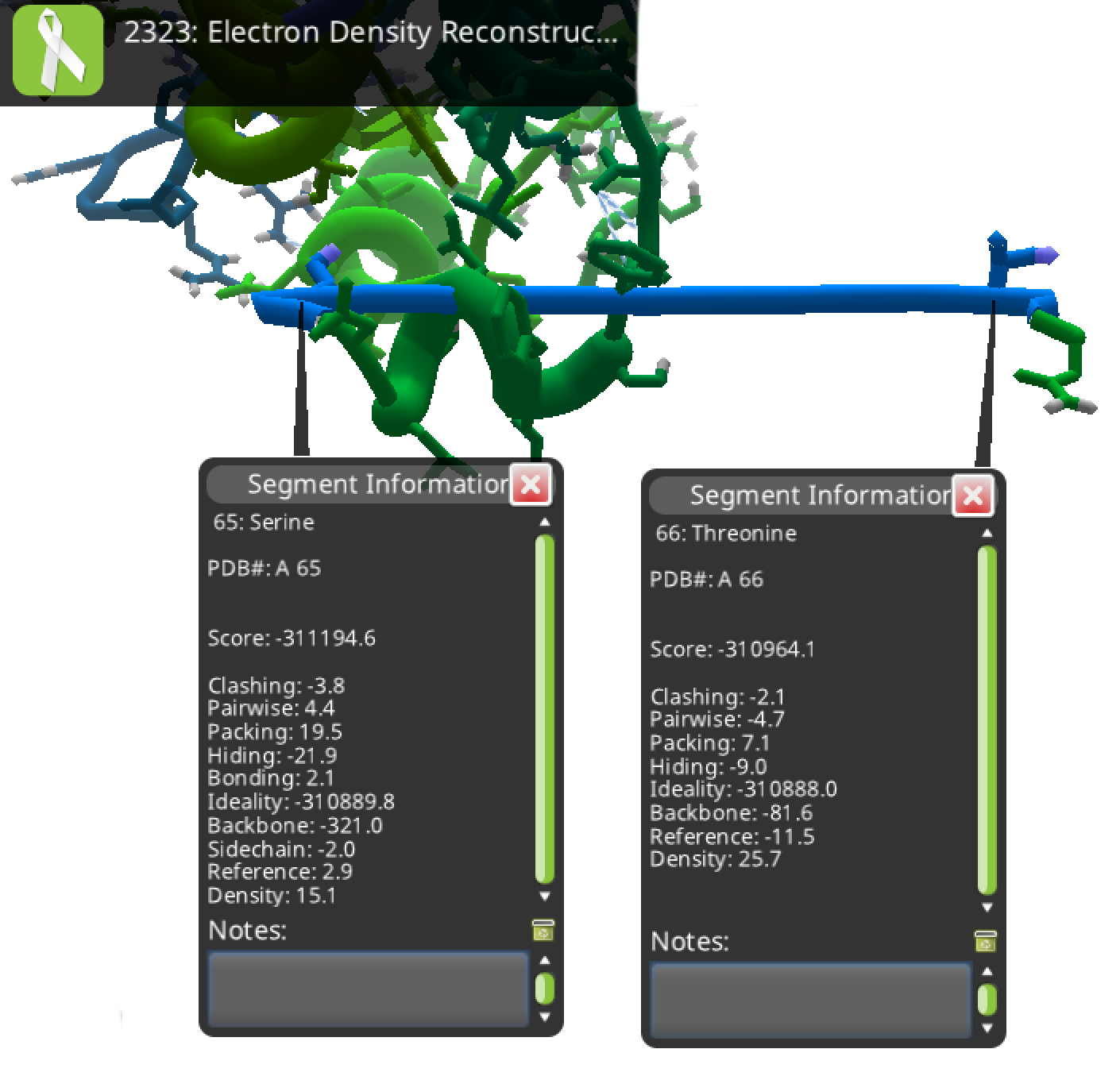
Foldit standalone produces the same structure from the 3HRY PDB file. Standalone also throws in the popup message "trunc variants detected" upon loading the PDB.
I used bands to measure the distances between the atoms of segment 65 and segment 66. The mean distance for successful bands was 27.33 Angstroms. For some reason, bands failed between atoms 1 and 4 of both segments, which I'm still looking into.
Since a PDB file doesn't usually include explicit bonds, it's up to the rendering software to figure out where the bonds should go. It seems like Foldit/Rosetta is doing this differently as compared to other viewers. Bonding atoms which are around 27 Angstroms apart doesn't seem reasonable.
PDB 3HRY does have a lot of oddities. There's a missing atom record for the final "OG" oxygen of residue A 65. Serine A 65 has multiple positions, so there are "ASER" and "BSER" records, but not for every atom. Similar "A" and "B" records appear in several other spots. Threonine A 66 has "B" records, but no "A" records.
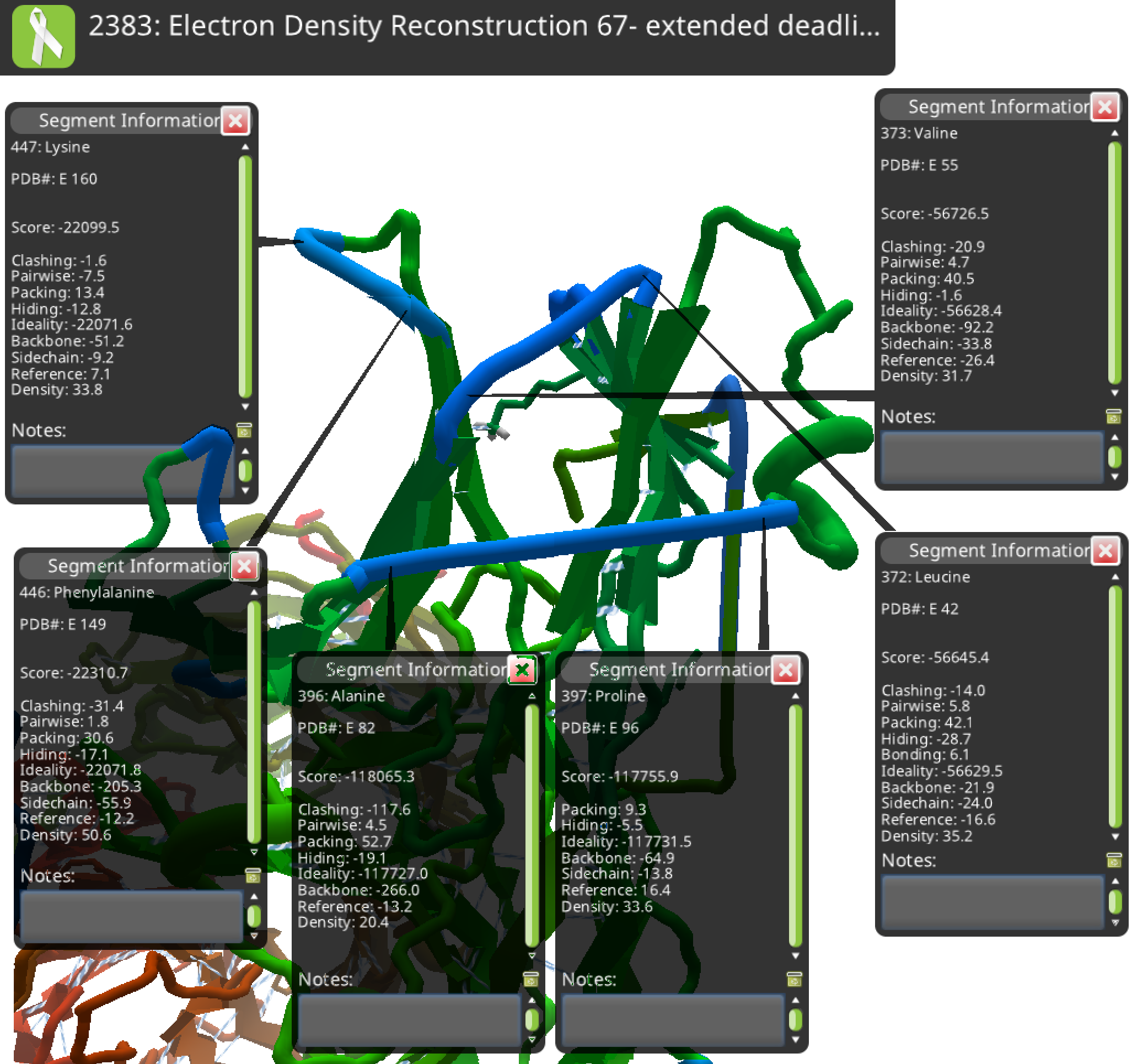
I'll look at 2383 next, which had lots of puckers and a few straws. So far, I think all those problems are due to missing residues.
I'll also look at some of the puzzles which ended up without puckers or straws.
See the bug report puzzle 2323 issues with missing densities for an earlier look at 2323.