In puzzle 2323, there is an artificial jump in the structure due to alternate conformations with different amount of visible residues. This could be misleading or causing some kind of inaccuracies in ED reconstruction.
I also assume that when cutting out the density around the protein, altloc B of residues 58-65 in chain A have been ignored, thus the densities are not showing in the game.
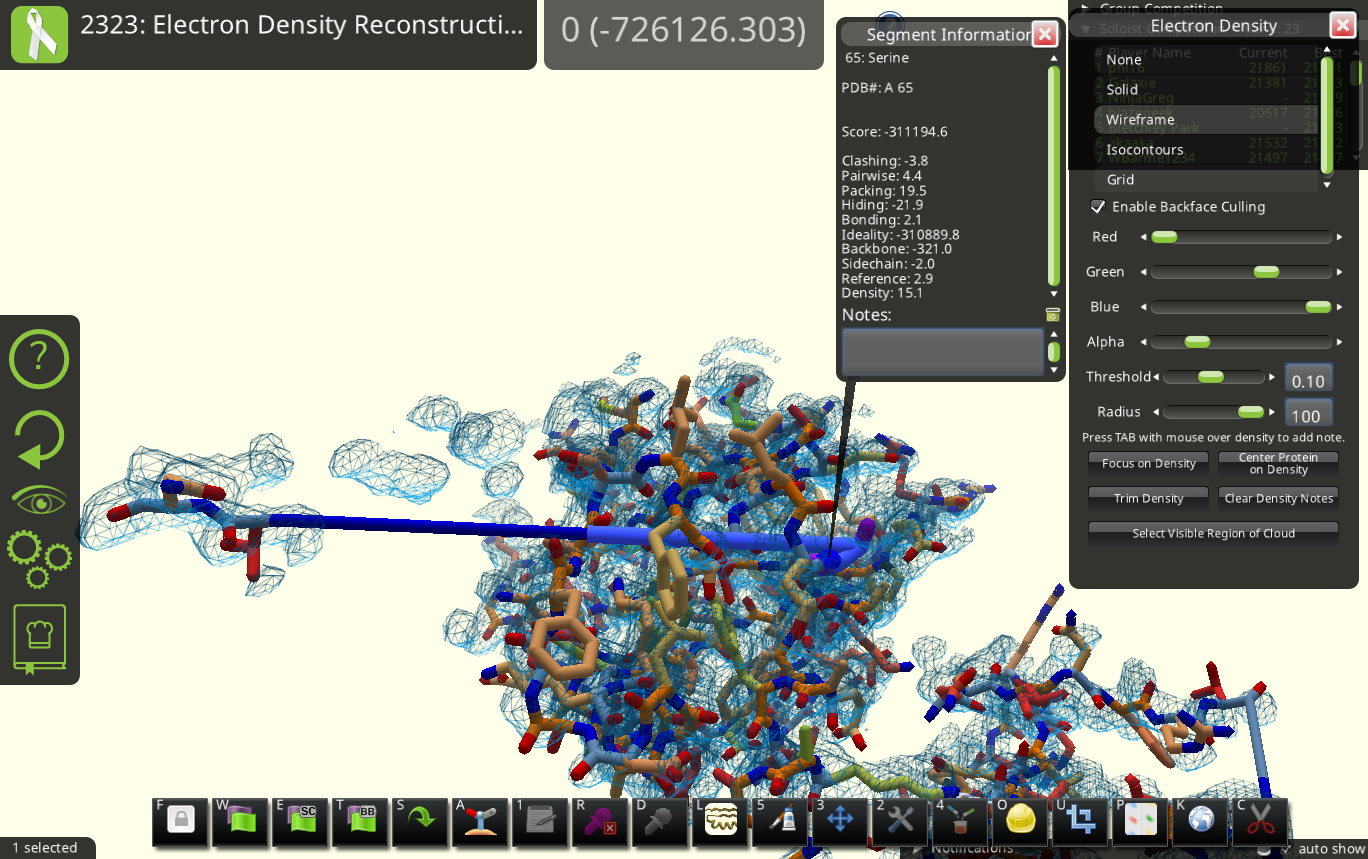 (edit: fix protein display)
(edit: fix protein display)
From a pdb search of the sequence, this structure should be chains A-C of PDB ID 3HRY (correct me if I was looking at the wrong PDB). In chain A, only alt A locations were provided for residues(segments) 58-65, while residues 66-67 is only visible in alt B. The two altloc at the point of "disconnection" (S65) are highlighted in pink in the following figure. (density contour at 1.0 sigma, map converted to ccp4 format with phenix, image rendered in pymol)
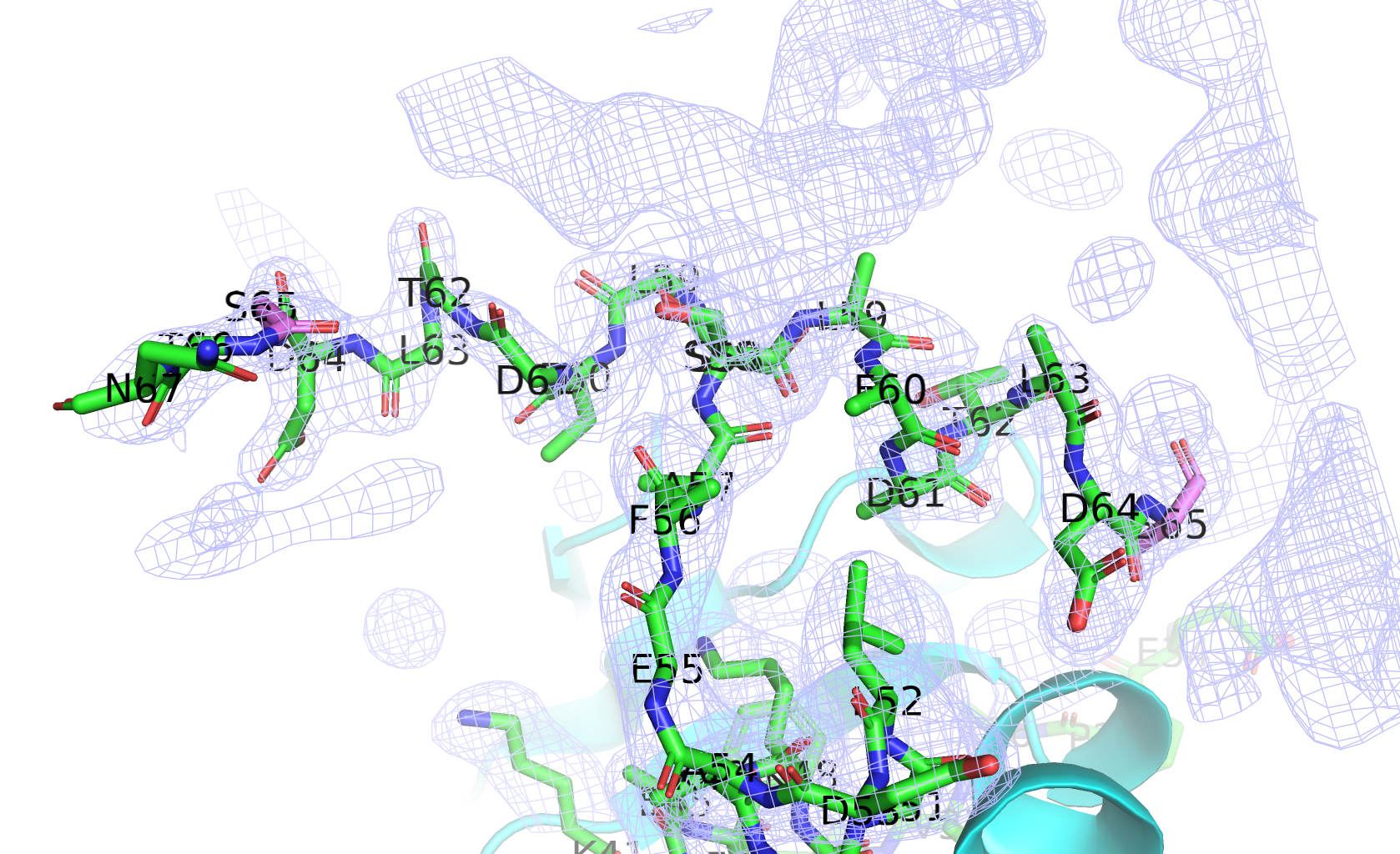
Another problem is in chain C, where residue 8 is connected to residue 14. There should be some gaps between.
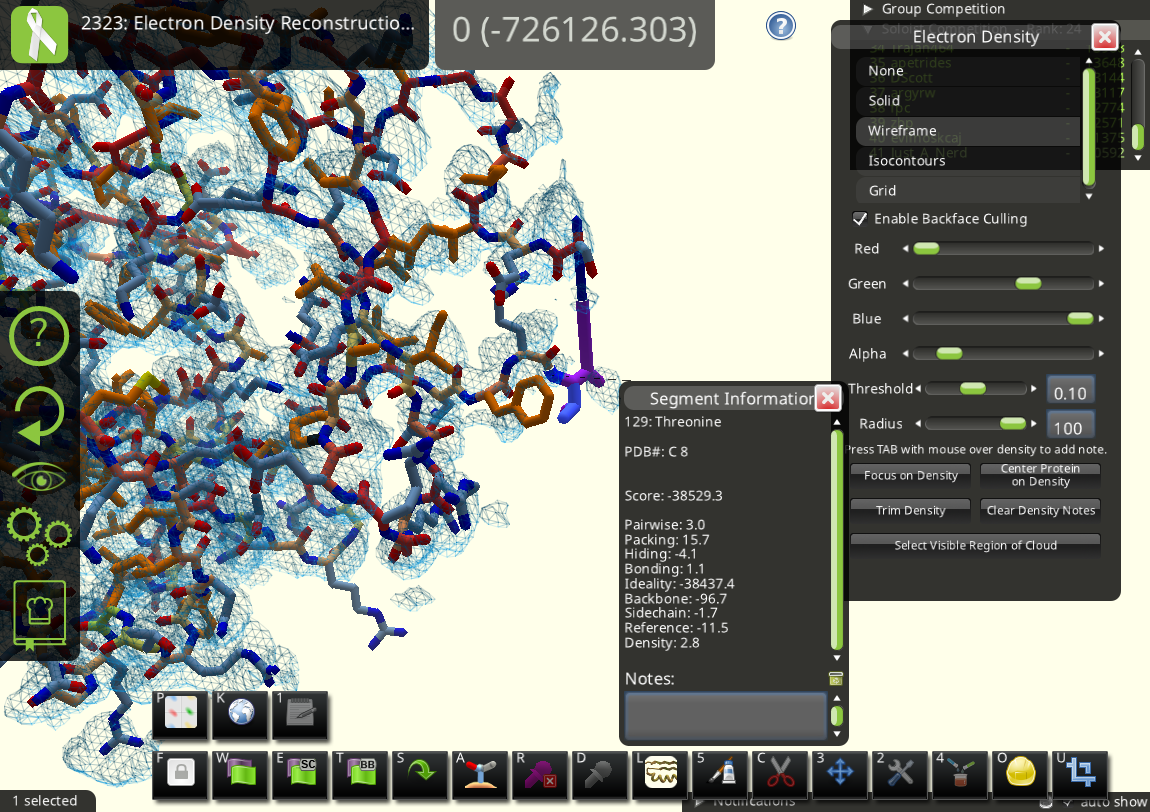
Since residues 9-13 are disordered and hence not visible in the X-ray density, a better way to handle this is probably separating the two segments 8 and 14 (highlighted in orange in the following figure) instead of connecting them.
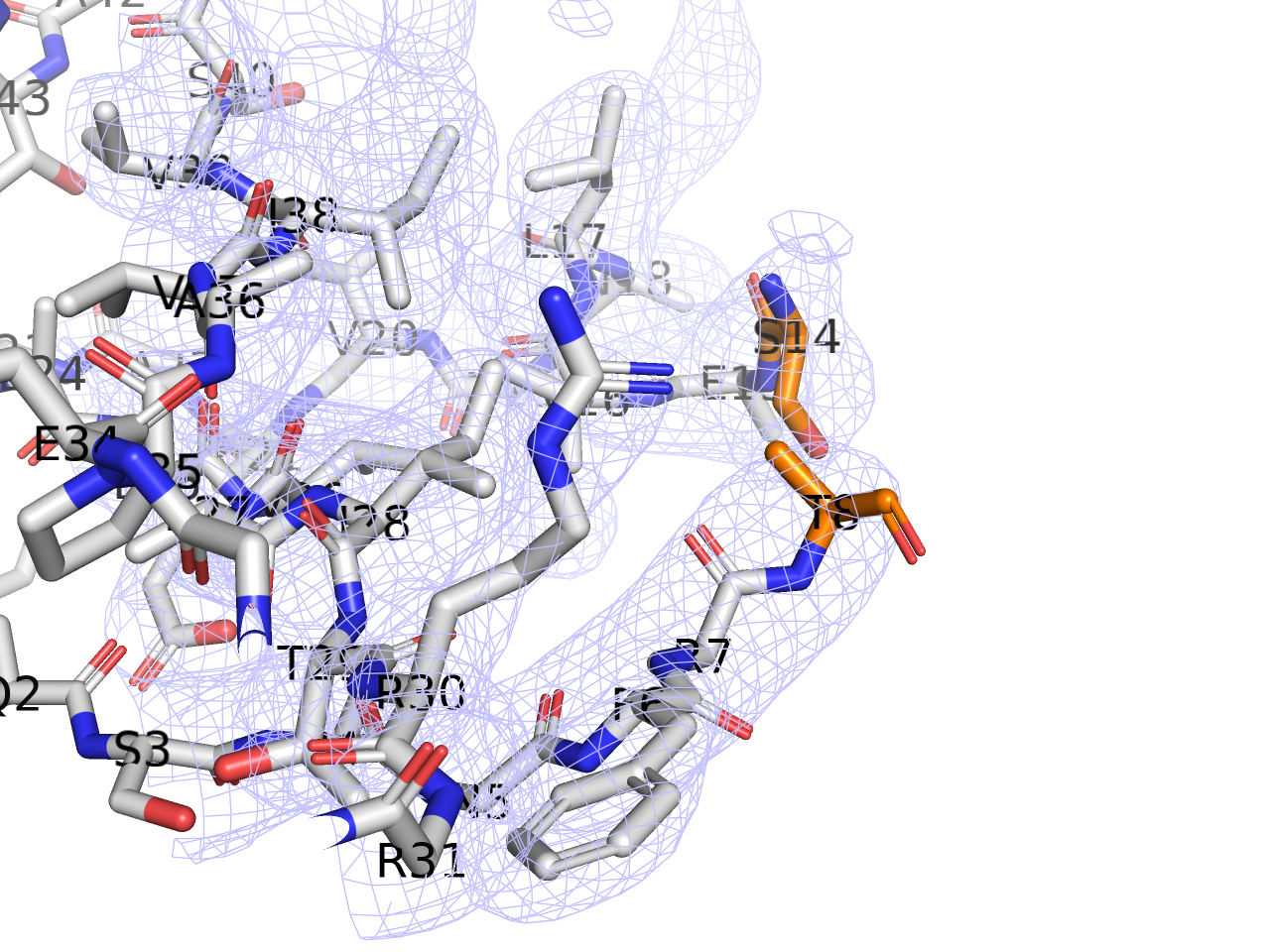
Out of curiosity, I tried to recalculate the map of 3HRY after removing alt. conf. B of residues 58-65 of chain A.
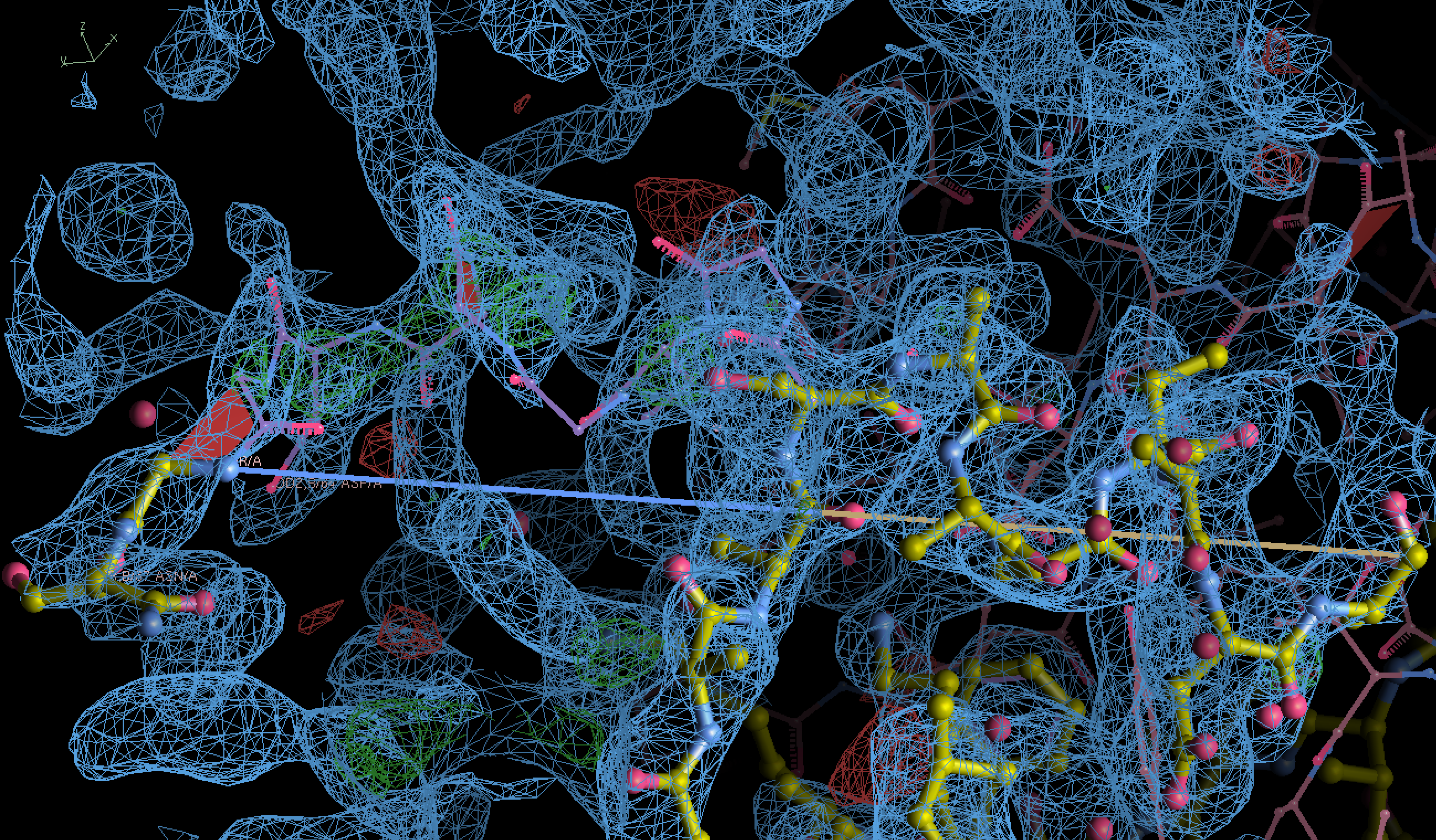
In the figure, 2Fo-Fc map is colored in blue and contoured at 1 r.m.s.d., and the Fo-Fc map colored in green/red for positive/negative densities contoured at +/- 3 r.m.s.d..
Model with deleted altloc displayed as yellow sticks, and the aligned original PDB with altloc B shown as purple wires.
There seems to be some green maps in the deleted regions, which could indicate unmodelled densities, of which the densities are missing in puzzle 2323.
Although the maps are much cleaner around the residues at alt. conf. A compared to alt. conf. B as I remove both altloc A and B of 59-67 in the model and recalculate the map in another trial (omit densities shown in the following figure)
Using the same contour level, but the deleted resides from alignment are shown as green ball-and-sticks for clarity, and the model displayed as wires.
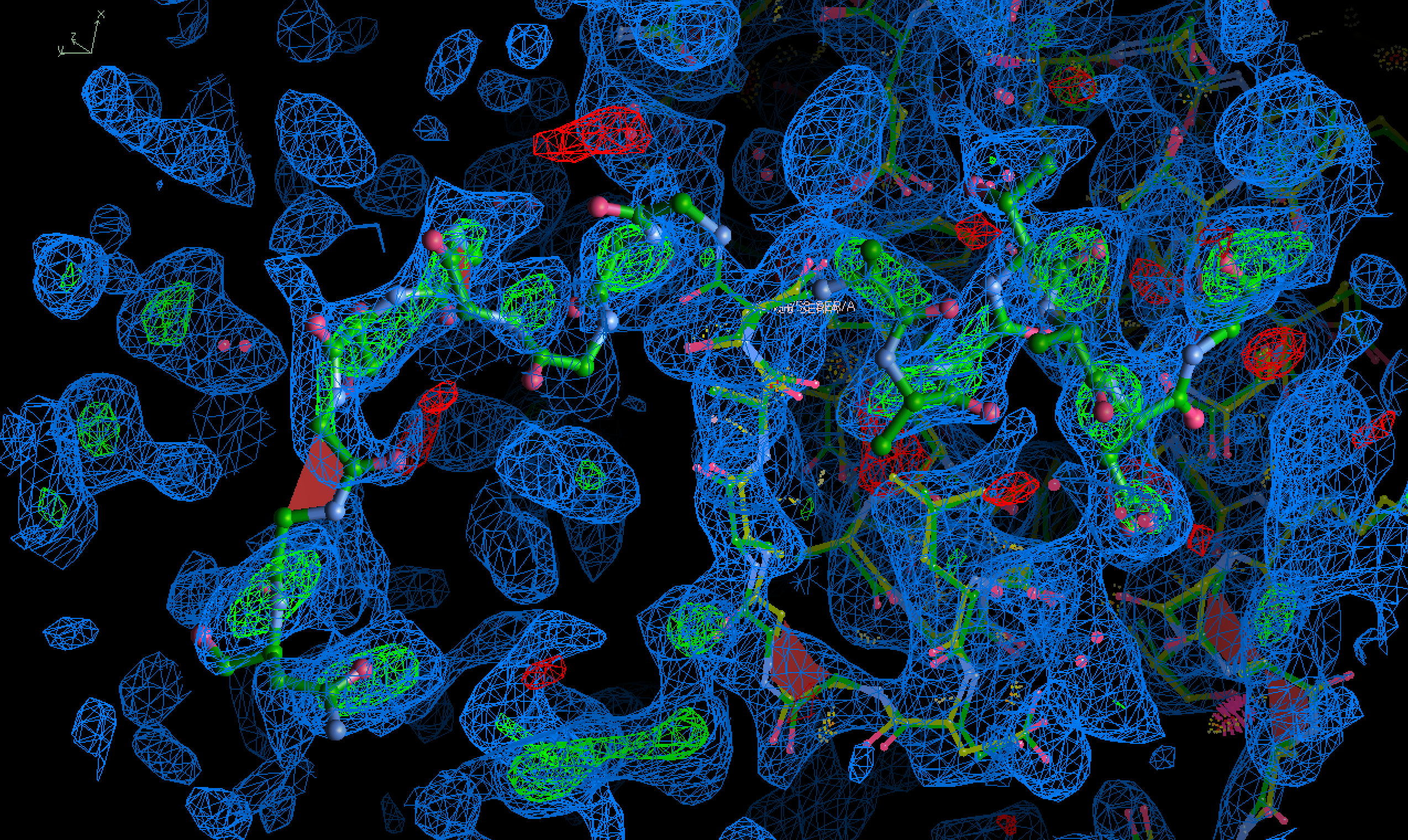
(recalculate map with phenix after deleting the selected residues, images rendered in WinCoot.)
(edit: added another trial for fun. Indeed the density view and the simplified controls in Foldit are more attractive for teaching purpose!)
Thanks- I agree it looks like this one might need some more hands-on work in a repost, but we might have some fun new ways of handling that soon…
Pz #2341 also has one (Q) and two residue (HQ) jumps on some un-modeled residues. At 1 rmsd for blue (using PDB-REDO density), the two ends for chain A appear to barely bridge together. Forcing WinCoot to fit loop (first add residue, then fit loop > rama search on sequence (I can't figure out how the database tool works)) seems to give something on the missing Q on chain A by pointing it towards a green blob, but I'm not very convinced that's the real stuff.
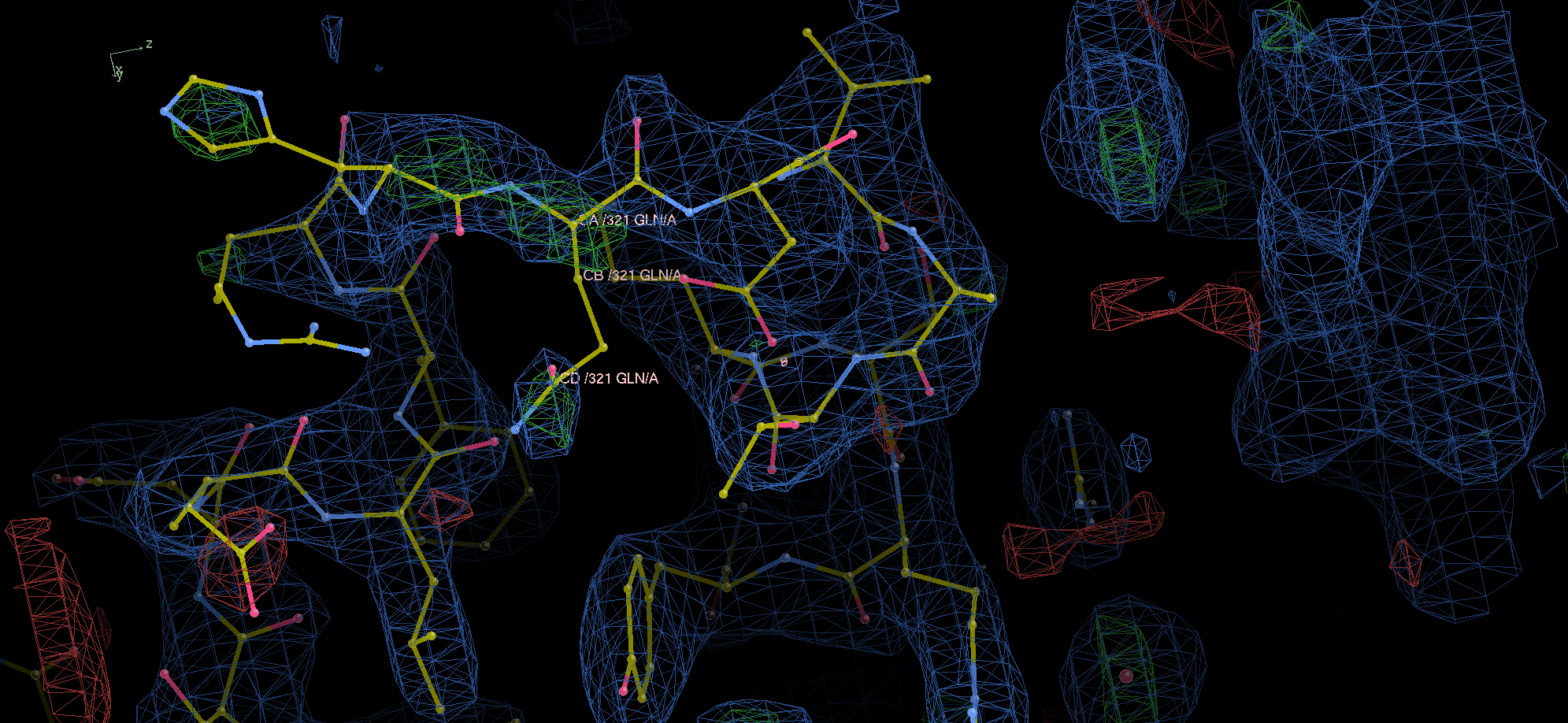
flipped some things
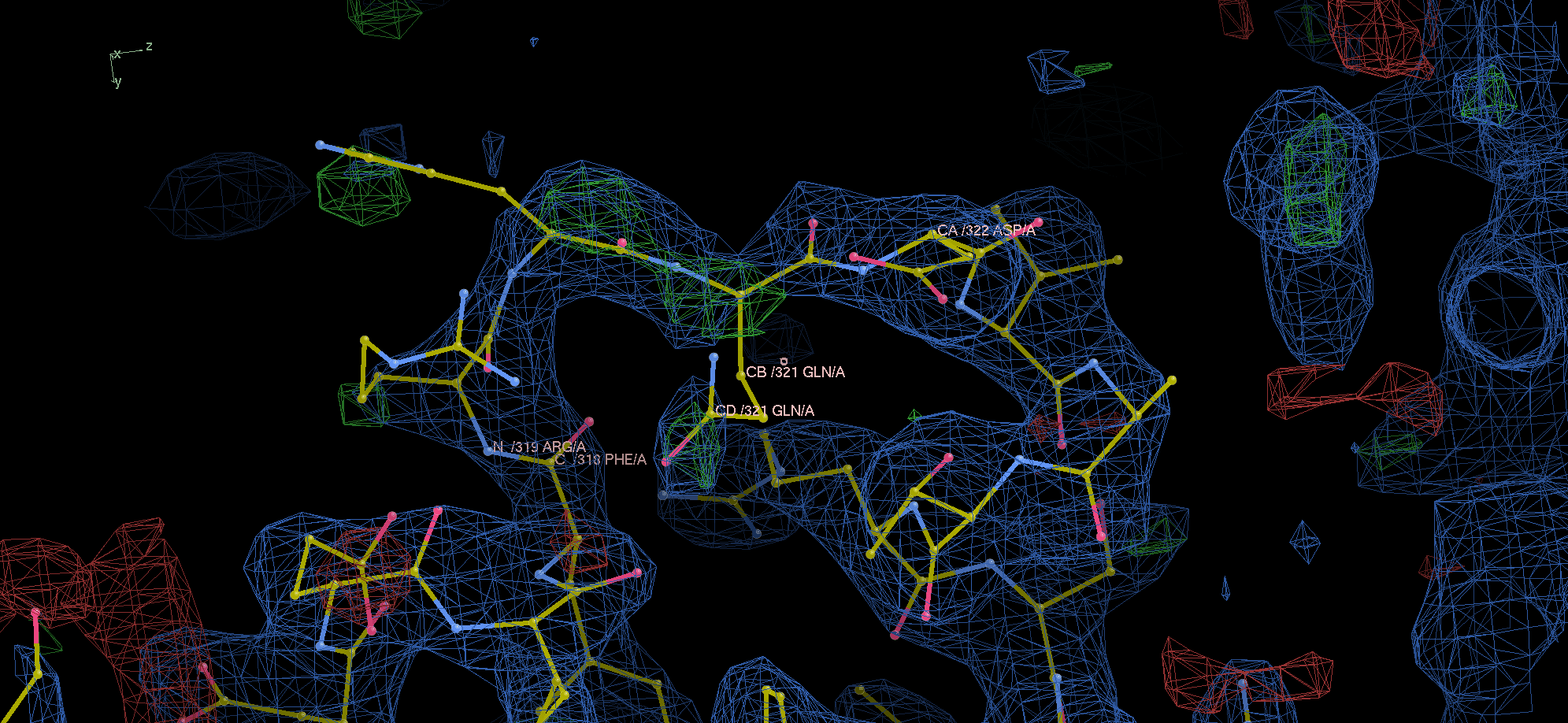
Chain B gap really has no density.
Thank you for bringing this up… our apologies that it took so long to respond (unfortunately this occurred over the Thanksgiving holidays).
We are looking into preventing this from happening again… thanks for your patience!
Way late, I'm now using the term pucker for the problems seen in 2383 and 2323. The odd part is that many ED recon puzzles handle missing residues in a reasonable way, with the sides of the gap ending up as separate chains. That's not perfect, but it's better than joining up the segments on either side of the gap as if nothing happened.
Edit: on further consideration, I've added the term straw for long straight sections with terrible ideality. Puzzle 2323 has a straw at 65-66, but it's not due to missing residues. Puzzle 2383 has straws and puckers due to missing residues, although I haven't finished a deep dive into the PDB file for that one.

 (edit: fix protein display)
(edit: fix protein display)








