
rosie4loop Lv 1
Motivation
Rational design of targeted covalent inhibitor or mechanistic probes are getting more popular nowadays. However, if we want to design non-covalent binders the functional groups involving in covalent modification (aka covalent warheads) should be avoided.
I was playing with a series of covalent warheads to check the value "bad group" penalties of the functional groups. Then I realized that they're placed near a random serine or cysteine. So I've decided to test if it's possible to do something similar to covalent docking in Foldit.
Workflow
- attach a covalent warhead onto the ligand
- wiggle to relax the conformation before it react
- modify the part of ligand that would be changed after reaction
- add a band to fake the covalent bond with protein
- changing CI, wiggle and shake to pull the bonded atoms together
- with some restraints(BiS of length 0) or freezes of bonded part, restore CI and relax other parts of the complex.
General comments
- the most important features lacking in Foldit for this purpose is interactively changing the clash importance of individual atoms.
- it'd be better if finer control of protonation can be done
- it's also better to have a torsional restraint/bonus and a modified scoring function for this purpose
Test using Puzzle 2307
Remarks: depends on the protein, covalent inhibition may or may not be practical/useful. I'm using this puzzle just to test the potential of foldit
- Model a non-covalent pose of a sulfonyl fluoride
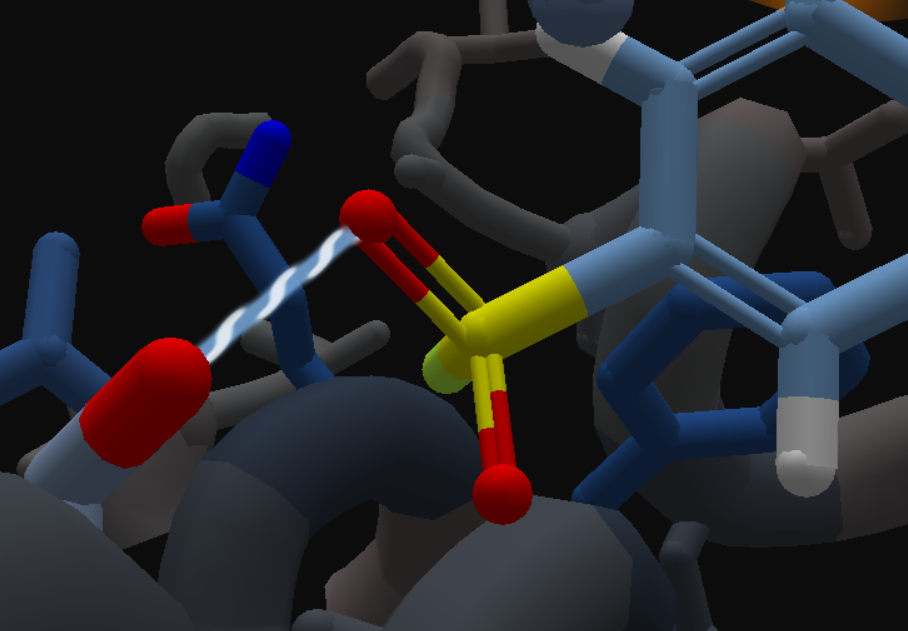
- Removing the fluorine atom
- Add a band to connect the serine @og with sulfur.
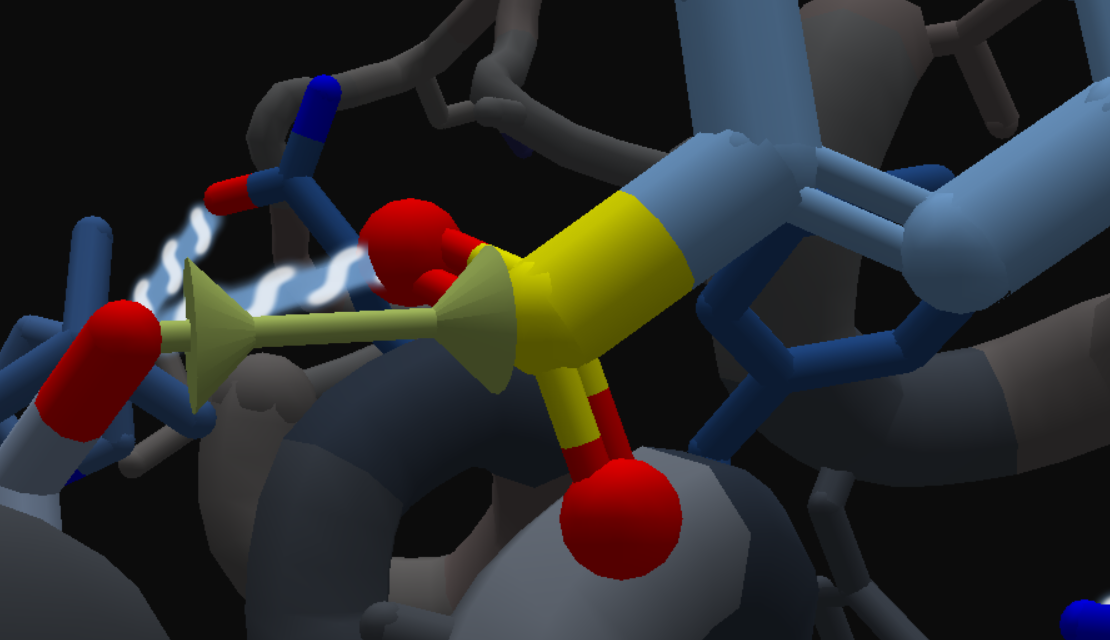
- Change CI.
- Relax the structure with shake and wiggle to pull the "bonded atoms" together. This part is difficult with the current Foldit, it's difficult to proceed from here. Even if I turn the H-bonds importance to zero, oxygen in the sulfonyl group tends to move towards the serine and push the sulfur away.
Unsuccessful attempt so far, but the potential is there if the items in general comments can be implemented.