1841b: Coronavirus NSP6 Prediction
Closed since almost 6 years ago
Intermediate Intermediate Intermediate Intermediate Intermediate Intermediate Overall Overall Overall Overall Overall Overall Prediction Prediction Prediction Prediction Prediction PredictionSummary
- Created
- May 22, 2020
- Expires
- Max points
- 100
Note: This puzzle replaces Puzzle 1841, which was accidentally posted with an incorrect sequence.
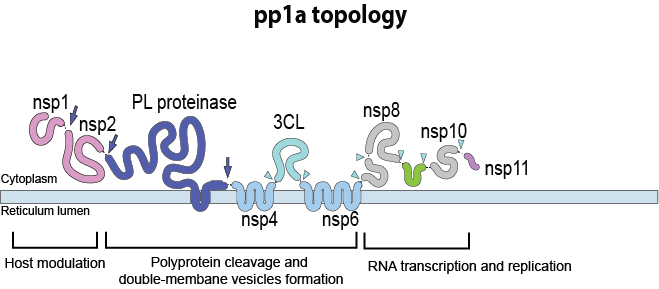
Fold this coronavirus protein! This is a portion of a larger protein encoded in the viral genome of SARS-CoV-2. It is encoded in a region of the genome called NSP6, but the protein's structure and function are still unknown. If we knew how this protein folds, we might be able to figure out its exact function. The puzzle's starting structure shows SS predictions from PSIPRED, and hints which parts of the protein might fold into helices or sheets. Refold this protein to find high-scoring solutions, which will tell us how this protein is most likely to fold!
Sequence:
CTCYFGLFCLLNRYFRLTLGVYDYLVSTQEFRYMNSQGLLPPKNSIDAFKLNIKLLGVGGKPCIKVATVQ
Top groups
-
1. Go Science100 pts. 9,500
-
-
-
-
-
-
-
-
-
Comments


puxatudo Lv 1
I agree with you Jeff101. The only thing I'm think is, if it's a transmembrane protein the scoring system should be more complex than that.
Considering the different types of membrane proteins (peripheral and integral, and their different relations to the bilayer), the scoring system couldn't be onefold (pun not intended! haha).
There should be two scoring systems, one for the part embedded in the membrane (for the intrinsic part of an integral membrane , let's say), and another for the extrinsic part of the protein.
You should get a high score if the hidrophobics would be sticking out for the INtrinsic part of the protein.
You should get a high score if the hidrophilics would be sticking out for the EXtrinsic part of the protein.
So, two scoring systems.
Or they could try to lock the extrinsic part and score the intrinsic, and vice-versa. Using two different scoring objectives.

Susume Lv 1
This is an article about the original SARS virus (2008), but it seems reasonable to speculate that the current virus behaves similarly: it invades the ER membrane and restructures it, using the resulting network of altered ER membrane as a place to replicate:
https://journals.plos.org/plosbiology/article?id=10.1371/journal.pbio.0060226
Here is a more recent summary (2015) of ways that positive RNA viruses use the ER membrane to form protective containers in which to synthesize, either by intruding or by extruding. When they extrude the membrane, they form DMV (double membrane vesicles), which are like protective bubbles isolated from the cytoplasm where their RNA can be synthesized without being attacked.
https://pubmed.ncbi.nlm.nih.gov/25287059/
So some of the coronavirus proteins must be involved in attacking and restructuring the ER membrane, and possibly also as structural elements of the resulting altered membrane and the DMVs.

bkoep Staff Lv 1
The NSP6 protein definitely associates with the membrane, but we think that this portion might form a well-folded domain in the cytosolic region of the cell.
This is far from proven, but there is some evidence to support it. This study from 2009 (working with a related coronavirus) used some cool techniques to study the orientation of the full protein in the membrane:
https://pubmed.ncbi.nlm.nih.gov/19386712/
It is still possible that this portion of the protein interacts with the membrane. But in this puzzle, we would like to predict how this domain might fold up if it were cytosolic.
We could consider a follow-up puzzle with a modified "membrane" score function, but Foldit is not very well suited to model the membrane environment. (As puxatudo points out, things get complicated.)

bkoep Staff Lv 1
We use our standard score function for these puzzles.
This is potentially a problem, if this domain makes significant contacts with the rest of the protein, or with the membrane. Those contacts would not be scored appropriately with our standard score function, and could be problematic for predicting how the domain will fold.
However, many natural proteins consist of smaller domains that are perfectly capable of folding independently. If that's the case, then we should be able to use our standard score function.