Bruno Kestemont Lv 1
This remembers me a smaler de novo symetric puzzle (pentamer I think) that I designed several years ago with the same concept ;)


Closed since about 3 years ago
Novice Novice Novice Novice Novice Novice Overall Overall Overall Overall Overall Overall Prediction Prediction Prediction Prediction Prediction Prediction Electron Density Electron Density Electron Density Electron Density Electron Density Electron DensityThe structure of this protein has already been solved and published, but close inspection suggests that there are some problems with the published solution. We'd like to see if Foldit players can use the same electron density data to reconstruct a better model. There's three copies of the same protein here, but not all the segments are visible. It's pretty big, so this is one where the Trim tool might come in handy. One other note: if you happen to glance at the PDB entry; you might recognize the names of some of the authors for their involvement in Foldit. Not everything is perfect the first time around...

This remembers me a smaler de novo symetric puzzle (pentamer I think) that I designed several years ago with the same concept ;)

Is this a called a symmetric trimmer ? It seems that the 3 monomers are independent.

Yes, this one has three chains, 160 residues each. The chains are not 100% identical, so it may not be considered a true symmetry puzzle.
It's a match for 4KYZ and 4KY3, both mentioning "designed protein OR327".
For some reason, the sequence for 4KYZ shows four identical chains. The 3D viewer on rcsb.org only shows chain A. I'm not sure if there's an option to see all the chains.
When I open 4KYZ in Jmol, I see four chains, but they're in a totally different configuration than what we're seeing in 2282. This seems to be a versatile protein, sometimes a trimer, sometimes a tetramer.
Here are the chains detected by the latest version of "print protein", which I hope to release soon:
A: diqvqvniddngknfdytytvtteselqkvlnelmdyikkqgakrvrisitarsskeaykflailakvfaelgyndinrkmtvrfrgddlealekalkemirqarkfagtvtytldgndleititgvprqvleelakeaerlakefnitititvtvegql
B: iqvqvniddngknfdytytvtteselqkvlnelmdyikkqgakrvrisitarsskeaykflailakvfaelgyndinrkmtvrfrgddlealekalkemirqarkfagtvtytldgndleititgvprqvleelakeaerlakefnitititvtvegqlg
C: diqvqvniddngknfdytytvtteselqkvlnelmdyikkqgakrvrisitarsskeaykflailakvfaelgyndinrkmtvrfrgddlealekalkemirqarkfagtvtytldgndleititgvprqvleelakeaerlakefnitititvtvegql
This version of "print protein" relies on the distance between alpha carbons (from structure.GetDistance) to detect where chains begin and end. That's useful when atom counts don't tell the whole story.

I just noticed the "not all the segments are visible" comment in the official notes.
Each of the three chains is missing the MG at the beginning, and then the expression tag, GSLEHHHHHH at the end.
However, chain B is also missing the D at the beginning, and then picks up a G from the expression tag at the end, so the length is still 160.
The PDB file for 4KYZ doesn't really explain all this too well, but it does have "REMARK 465" records for some of the missing residues.
Another interesting point is the PDB file calls for selenomethionine (MSE) instead of plain methionine (MET). Selenomethionine substitutes a selenium atom in the place of the sulfur atom found in methionine. Foldit apparently doesn't work with selenomethionine, so we're stuck with plain old methionine.
Here's the Foldit sequence, in lower case, aligned with the stated sequence in upper case, with everything split into chains:
1 1 1 1 1 1 1
1 2 3 4 5 6 7 8 9 0 1 2 3 4 5 6
1234567890123456789012345678901234567890123456789012345678901234567890123456789012345678901234567890123456789012345678901234567890123456789012345678901234567890
diqvqvniddngknfdytytvtteselqkvlnelmdyikkqgakrvrisitarsskeaykflailakvfaelgyndinrkmtvrfrgddlealekalkemirqarkfagtvtytldgndleititgvprqvleelakeaerlakefnitititvtvegql
MGDIQVQVNIDDNGKNFDYTYTVTTESELQKVLNELMDYIKKQGAKRVRISITARSSKEAYKFLAILAKVFAELGYNDINRKMTVRFRGDDLEALEKALKEMIRQARKFAGTVTYTLDGNDLEITITGVPRQVLEELAKEAERLAKEFNITITITVTVEGQLGSLEHHHHHH
iqvqvniddngknfdytytvtteselqkvlnelmdyikkqgakrvrisitarsskeaykflailakvfaelgyndinrkmtvrfrgddlealekalkemirqarkfagtvtytldgndleititgvprqvleelakeaerlakefnitititvtvegqlg
MGDIQVQVNIDDNGKNFDYTYTVTTESELQKVLNELMDYIKKQGAKRVRISITARSSKEAYKFLAILAKVFAELGYNDINRKMTVRFRGDDLEALEKALKEMIRQARKFAGTVTYTLDGNDLEITITGVPRQVLEELAKEAERLAKEFNITITITVTVEGQLGSLEHHHHHH
diqvqvniddngknfdytytvtteselqkvlnelmdyikkqgakrvrisitarsskeaykflailakvfaelgyndinrkmtvrfrgddlealekalkemirqarkfagtvtytldgndleititgvprqvleelakeaerlakefnitititvtvegql
MGDIQVQVNIDDNGKNFDYTYTVTTESELQKVLNELMDYIKKQGAKRVRISITARSSKEAYKFLAILAKVFAELGYNDINRKMTVRFRGDDLEALEKALKEMIRQARKFAGTVTYTLDGNDLEITITGVPRQVLEELAKEAERLAKEFNITITITVTVEGQLGSLEHHHHHH

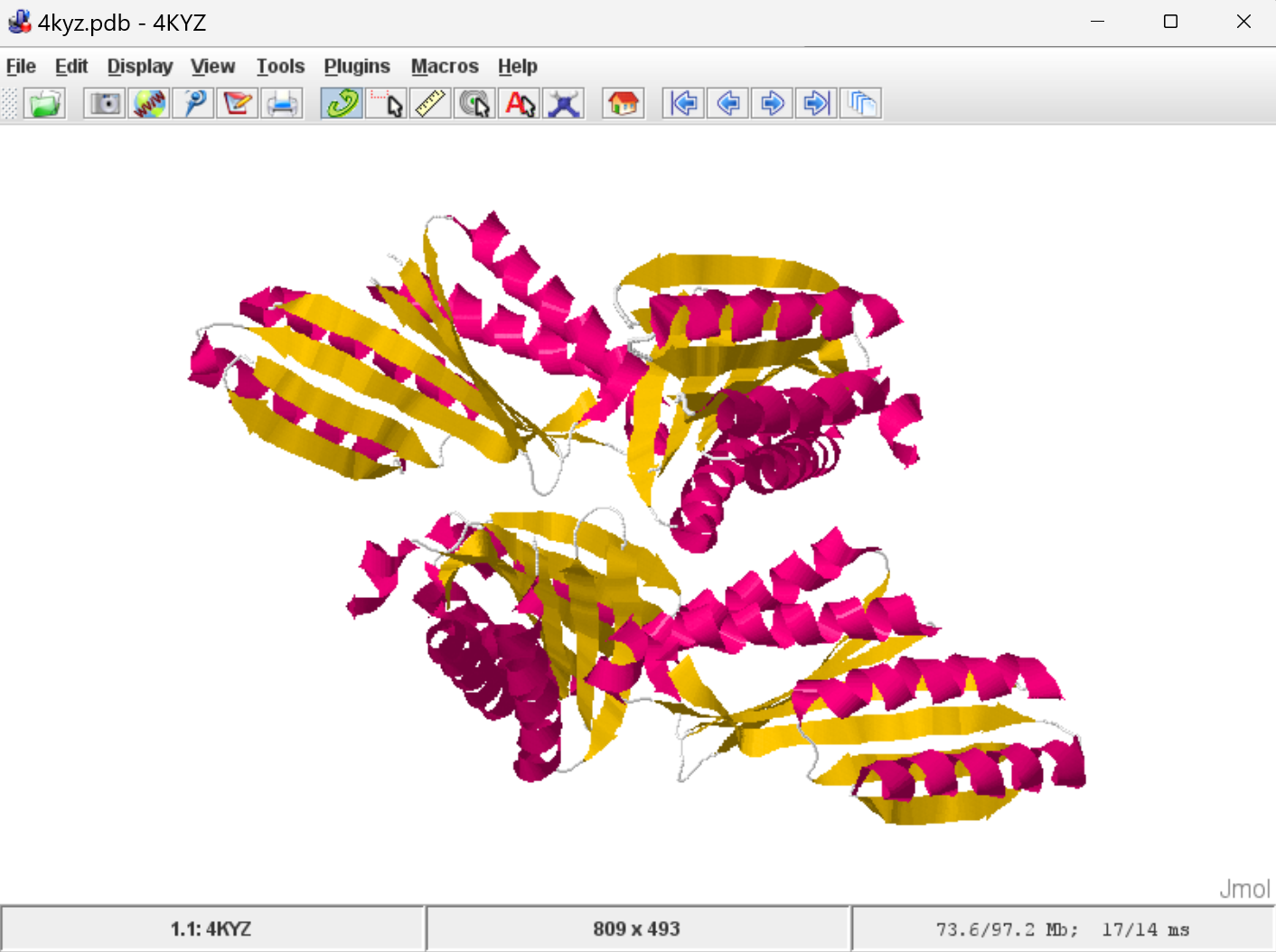
Here's what 4KYZ looks like: