LociOiling Lv 1
This week's puzzle includes DNA, and the protein part is pucker-free.
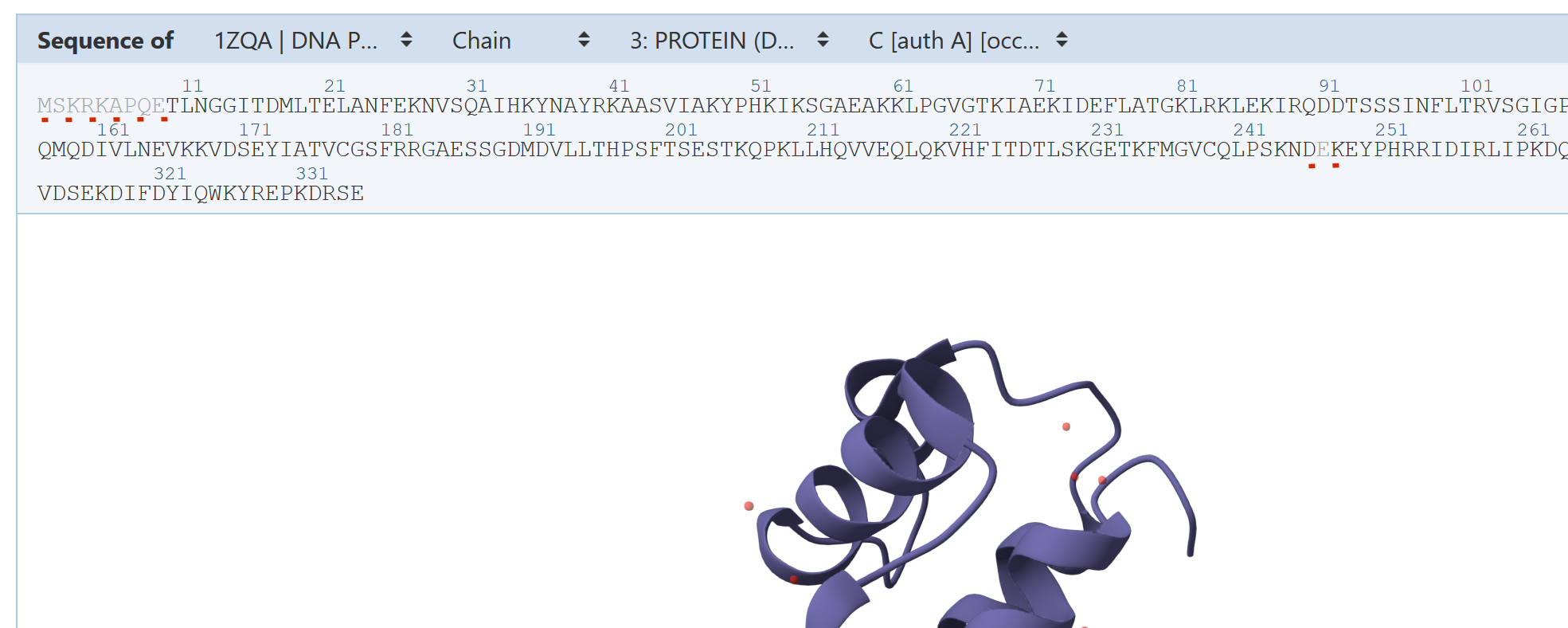
There are lots of matches for this protein, a DNA polymerase. There's a long series starting with 1ZQA and stretching to 1ZQT, which match both the DNA and the protein parts.
Unfortunately, at quick glance, all the entries in that range seem to start with "TLNGG", where the puzzle protein has begins "ETLNGG". So it's again a question of missing residues, those 1ZQ* entries seem to be missing one residue that the Foldit puzzle has, although I haven't inspected the entire list.
I will try to refine the search, the PDB assures me there are 678 matches for the protein part….