beta_helix Staff Lv 1
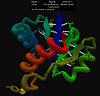
This is the page for this CASP10 refinement target:
http://predictioncenter.org/casp10/target.cgi?id=110&view=refinement
Note that the organizers give us this as a hint:
"N-term res. 1-4 are disordered in the native structure and removed from the starting model. Starting GDT_TS=84."
So, the first 4 residues have been deleted.
GDT_TS=84 essentially means that 84% of the model superimposes correctly onto the native structure:
We believe that Zhang-Server_TS4.pdb is identical to the refinement model (except that it obviously doesn't have the first 4 residues trimmed!)
Here is the sequence logo predicted by the SAM server:
H = helix
E = sheet
C = loop (or coil)
The taller the letter at each position, the higher the probability of that specific secondary structure for that amino acid.
For example, the Valines at residues 8, 11, 35-36, 62 & 69 are highly predicted to be part of a helix. However, the Valines at residues 49 & 52 are predicted to be in either loops or sheets.