beta_helix Staff Lv 1
Please look at the Notes for this puzzle (by pressing 3 in the Original Interface, or pressing 1 in the Selection Interface).
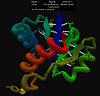
We recommend focusing your work on these 3 regions.
This is the page for this CASP10 refinement target:
http://predictioncenter.org/casp10/target.cgi?id=203
The CASP10 organizers gave us this hint:
"Starting model GDT_TS=90. Special attention regions: 41-50, 100-110, 125-128."
GDT_TS=90 essentially means that 90% of the model superimposes correctly onto the native structure:
Here is the sequence logo predicted by the SAM server:
H = helix
E = sheet
C = loop (or coil)
The taller the letter at each position, the higher the probability of that specific secondary structure for that amino acid.
For example, the Valines at residues 80-81, 94, 114-115, 123 & 134 are highly predicted to be part of a sheet. However, the Valines at residues 45, 52 & 103 are predicted to be in anything.