rosie4loop Lv 1
I am posting my notes here for those who would like to know the detailed steps to create custom Foldit puzzles with FolditStandalone and share it privately, e.g. for the training of students using in-house data yet to be published, or if you have concerns to let students create a game account.
You may also refer to the following article first for a brief overview of using Foldit in class:
- Official Foldit forum post for educators which provide detailed information on how to create the puzzle config. files (although lacking instructions on more updated tools or puzzle types)
- Dsilva, L., Mittal, S., Koepnick, B., Flatten, J., Cooper, S., and Horowitz, S. (2019) Creating custom Foldit puzzles for teaching biochemistry. Biochemistry and Molecular Biology Education 47, 133–139. [link]
Just to test the potential of custom Foldit puzzles for educational purposes. Please let me know for any issues – I am still new to Foldit and xtal structure solution.
Different concepts can be illustrated with the same puzzle, instructors should decide what to emphasize on according to the intended learning outcomes. For example,
- puzzle B let the student do something similar to what a crystallographer does to check how mutating the protein changes its conformation, ultizing the Foldit interface that is simpler than the actual tool
- combining puzzles B and C one could demonstrate how a point mutation in the active site results in the lost of drug affinity.
Hence I am skipping the aim of each puzzle in my discussion, focusing only on the technical details.
Please also note that I am only testing the minimal setup of puzzles, WITHOUT the “.puzzle_setup” configuration file that restricts movement, locks tools, limits range of mutatable residues etc. You may refer to the instructions provided by Foldit developers on how to create the setup files.
Disclaimer:
This is only a test on creating custom Foldit puzzles for educational purpose to demonstrate the process of manual model refinement, taking advantage of Foldit's comprehensive in-app biochemistry tutorials for an all-in-one learning experience. It should NOT be used for practical structure solution NOR as training material of structure solution from X-ray data.
This is not recommended for practical manual remodel since the 2Fo-Fc map is known to be model biased. Instead, use your favourite crystallography software like PHENIX/CCP4 and coot to (re)calculate the omit map of the poorly modelled region and model on that.
But I know that the 3D rendering in Foldit is attractive especially for teaching purpose, compared to the simple line-rendering in WinCoot/Coot (see the figure below!).
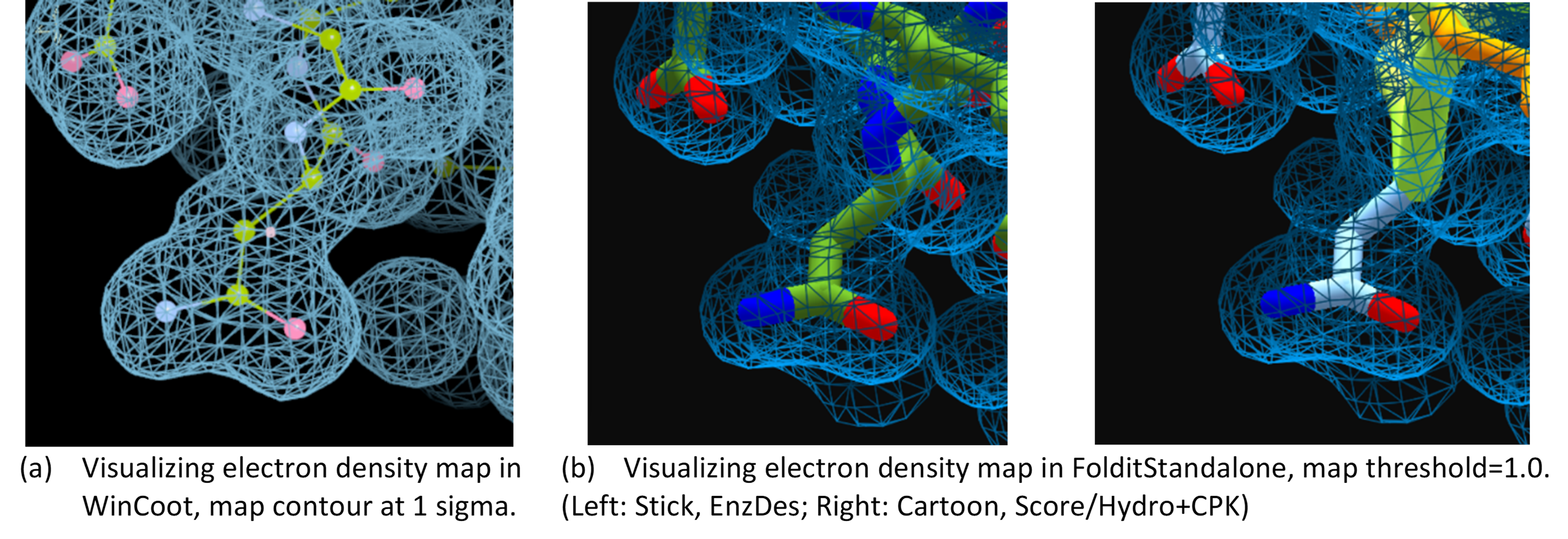
Please consult relevant materials, e.g. documentation and tutorials of PHENIX or CCP4 for methods of practical X-ray structure determination.
Contents:
A. A minimal, standard electron density puzzle
B. Electron density puzzle to model a mutant structure starting from a wildtype structure
C. ”Guess the ligand” electron density puzzle (TBD)
Software versions in this test:
- Phenix 1.20.1-4487 and WinCoot 0.9.8.1 on Windows 10,
- Folditstandalone: 20230109-b54c9a359c-win_x64-INTERNAL on Windows 10
- Rosetta 3.12 bundle compiled on Linux Mint 19.1
Other tools
• (Puzzle B) Your favourite tool of structure modelling and alignment, e.g. UCSF Chimera, PyMOL, TM-align, or can use the superposition tool in phenix as outlined here.
• (Puzzle C) Text editor like Notepad/Notepad++/gedit
Summary of the files required to set up each puzzle:
FolditStandalone input for puzzle A
- xxx.pdb
- Starting model
- xxx.density
- 2mfo-dfc map in ccp4 format
- xxx.wts_patch
- file telling Foldit how to score with the density info. If not included as you import the files, foldit would show you a warning.
ADDITIONAL input for puzzle B
- xxx.align
- Aligned fasta of template and query
- xxx.template.pdb
- Template structure pdb
ADDITIONAL input for puzzle C
- xxx.params
- Ligand params in rosetta format
Reference:
- Phenix documentation on map trimming and mtz2map
- Rosetta documentation about reading electron density map files
- FolditStandalone quickstart guide
- My previous notes on setting up a custom, minimal small-molecule design puzzle
- Dsilva, L., Mittal, S., Koepnick, B., Flatten, J., Cooper, S., and Horowitz, S. (2019) Creating custom Foldit puzzles for teaching biochemistry. Biochemistry and Molecular Biology Education 47, 133–139. [article][example files]
- Official Foldit forum post for educators, with the instructions and examples.