
rmoretti Staff Lv 1
We're excited to announce that Foldit players have been selected to participate in the next Critical Assessment of Computational Hit-Finding Experiments (CACHE) challenge.
CACHE
CACHE was conceived by the Structural Genomics Consortium and Target 2035 as the equivalent of CASP for small molecule drug design. (Original announcement paper in Nature Reviews Chemistry.) Teams from around the world are invited to design drug-like molecules which will bind to a particular target of interest. Each team will use their own computational methodology to determine the structure of ligands which not only bind well the to structure, but also have particular drug-like properties.
CACHE Challenge #2 – SARS-CoV-2 helicase
The next CACHE challenge is an exciting one – designing inhibitors for the SARS-CoV-2 helicase. The viral helicase is an important enzyme which is needed for restructuring the viral genome during replication. Inhibiting the helicase would greatly slow down viral growth, but right now there aren't any drugs targeting the helicase. Instead, the current approved vaccines induce your immune system to recognize the virus spike protein. Paxlovid is a combination of two inhibitors of the viral protease. Molnupiravir is an inhibitor against the viral RNA polymerase. Having an inhibitor for the helicase as well would open up the possibility of combination therapy approaches for COVID-19, an approach which has worked well in HIV treatment. Having multiple drugs works much better against rapidly mutating viruses, as it makes it much less likely that a virus would develop a variant which is resistant to all of the drugs.
The new Compound Library panel
In contrast to CASP, where the best answer already known, for CACHE the best answer is a wide open question. There are some structures of the helicase in combination with small molecules, but none which would make a good drug. The CACHE organizers will instead obtain and test submitted molecules. – This poses a challenge, as synthesizing novel molecules is a tricky and expensive prospect. There's a workaround, though. Recently there's been a growth of ultra-large make-on-demand compound libraries. These are libraries of a huge number (billions) of compounds which don't yet exist, but chemical suppliers think they can easily (and inexpensively) make. Using these ultra-large libraries means the CACHE organizers can test many more compounds than if they needed to be made from scratch.
To support these libraries, we've added a new tool to Foldit – the Compound Library panel.
![]()
As these libraries are too large to effectively search on the local client, clicking the Compound Library button will bring up an interface similar to the AlphaFold panel. This will allow you to submit your current small molecule designs to the Foldit server.
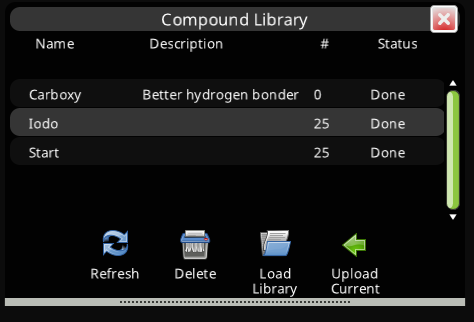
In the background, the Foldit server will search these ultra-large libraries. When the result is ready, the compounds in the library most similar to your design will show up in your Compound Library panel, where you can replace your current working ligand with the library compound.
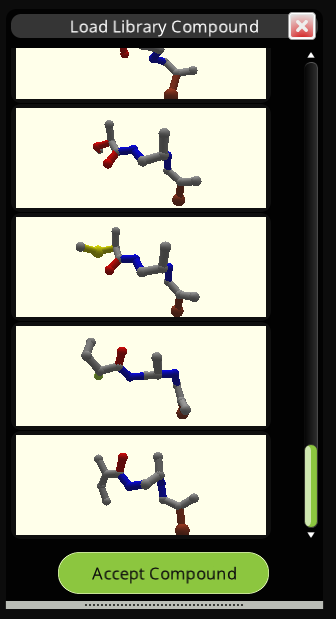
The concept is that you'll be able to go back and forth between the Small Molecule Design panel and the Compound Library panel. Use the Small Molecule Design tools to improve the binding and other properties of the ligand. Then use the Compound Library panel to find those molecules in the library which best match. You can then further modify and refine the library molecules with the Small Molecule Design tools, after which you can go back to the Compound Library to find improved molecules.
Foldit competes in CACHE!
We're planning to have a number of puzzle rounds between now and when the CACHE challenge closes for submission in late-November. From all of the puzzle results we will select the 100-150 compounds to submit to the CACHE organizers. Note that due to the limitations of synthesis, only compounds which are available in the library will be selected. So if you want your designs to be experimentally tested, it's best to work the Compound Library panel into your process. We will do similarity searches on top results, so they won't be completely ignored, but you'll likely get better results if you do the searching yourself.
Participation in CACHE puzzles is subject to the CACHE Terms of Participation, in particular “the Challenge IP [including Challenge Compounds] will be made freely available in the public domain pursuant to Creative Commons Attribution Only (CC-BY 4.0 or subsequent versions) licensing terms, with the intent that such Challenge IP may be Used and practiced by Users for any purpose”.
Special thanks to John Irwin and the ZINC database for the ultra-large library compound search behind the Compound Library panel.
P.S. Your previous efforts in small molecule design have not been forgotten! We are getting some exciting results from the VHL puzzle series, and we hope to bring you news on that front in the near future.

