Sometimes in the Ligand Design puzzles, I have tried to convert a high-scoring non-library compound into an on-library one. Generally, I lost points when doing so, but maybe it is worth sharing some tricks I have learned in the process.
Below is a sample compound that I drew in the ZINC20 database at https://zinc20.docking.org/substances/home/:
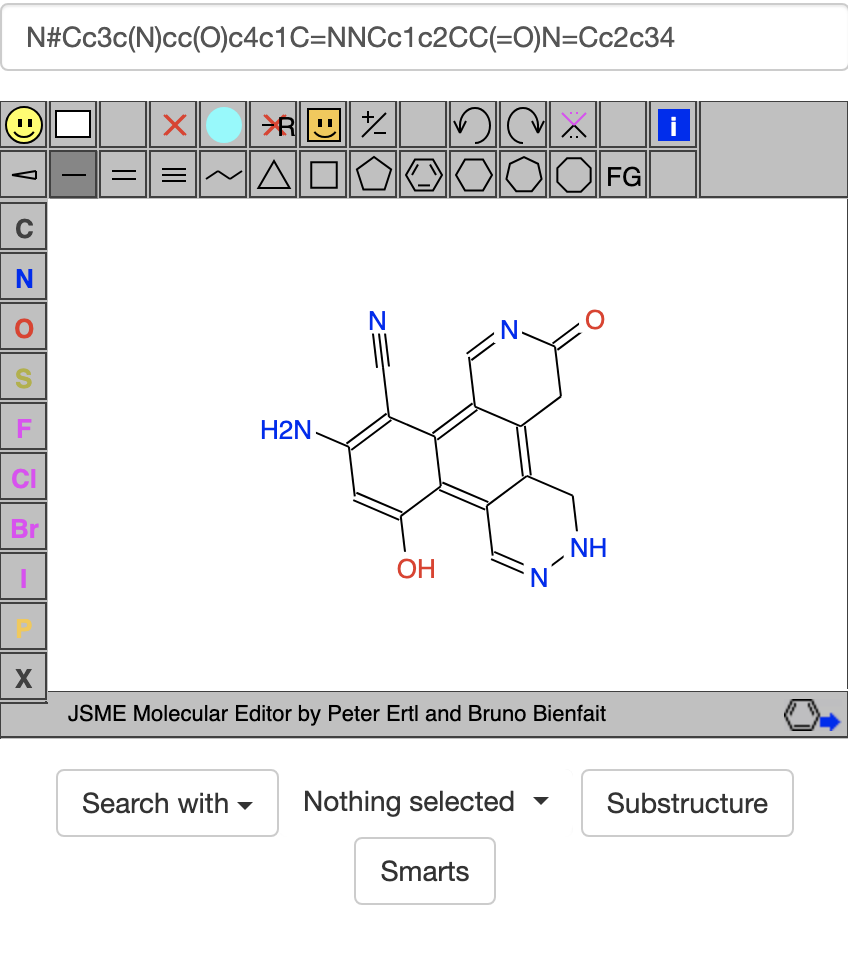
It has the SMILES code N#Cc3c(N)cc(O)c4c1C=NNCc1c2CC(=O)N=Cc2c34. If I press enter in the box containing the SMILES code, it searches for this compound in the ZINC20 library and says this compound has mol wt 305.297, logP 0.938, and 23 heavy atoms but is not in the ZINC20 library. Usually if I find something that exists in the ZINC20 library, there is a good chance that it will at least give hits (if not exact matches) in Foldit's Compound Library. Nevertheless, in Puzzles like 2404, I got the sense that compounds with 36 or more heavy atoms or molecular weight of 496 or larger were just too big to give hits in Foldit's Compound Library.
The compound above has 4 rings, 7 acceptors, and depending how you count things, 3-4 donors. At some point I tried https://zinc20.docking.org/rings/?ringcount=4&hba=7&hbd=3, which gave many interesting-looking compounds. One that stood out to me had 1 hexagon with 3 pentagons around it and the SMILES code C1=NNC2C3NN=CN3c3[nH]ccc3N12: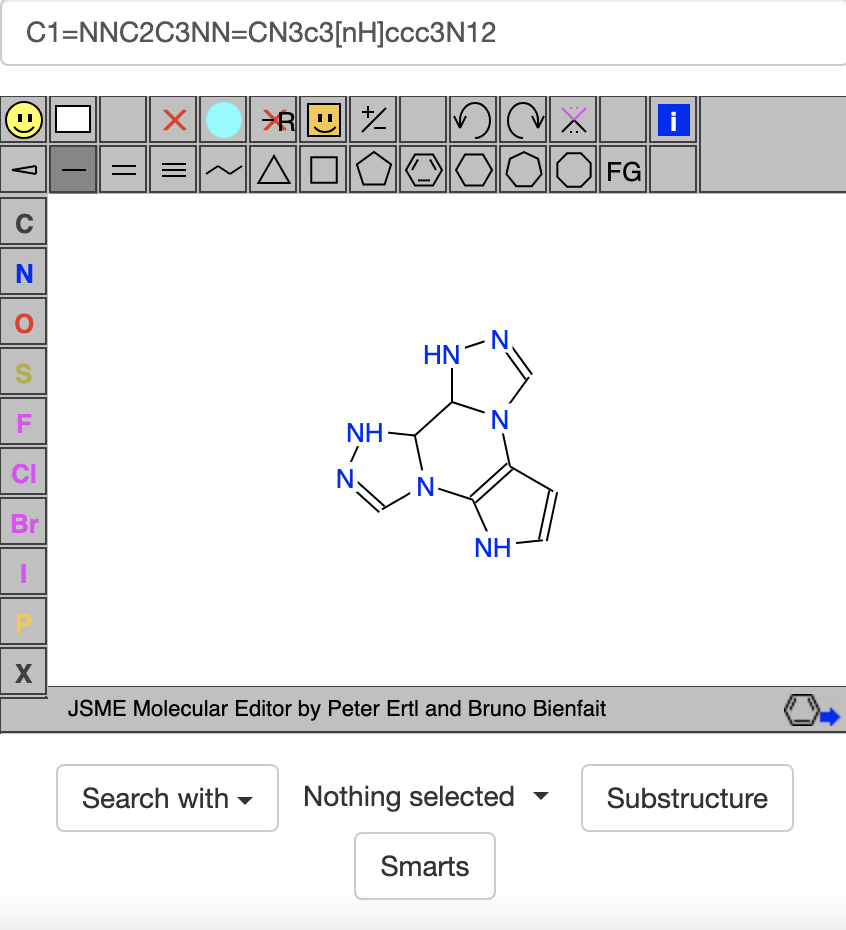
I also tried https://zinc20.docking.org/rings/?ringcount=4&hba=7&hbd=4, but on the first page of results, I didn't see any that looked as good to me as C1=NNC2C3NN=CN3c3[nH]ccc3N12 above did.
To find other compounds like this, I replaced each C c and N in the SMILES code with an *. I also replaced [nH] with an * and removed the = symbols for double-bonds. * is a wild-card character in SMARTS that can stand for any atom. I ended with the SMARTS code *1***2*3****3*3****3*12 which gave in https://zinc20.docking.org/substances/home/ the image below: 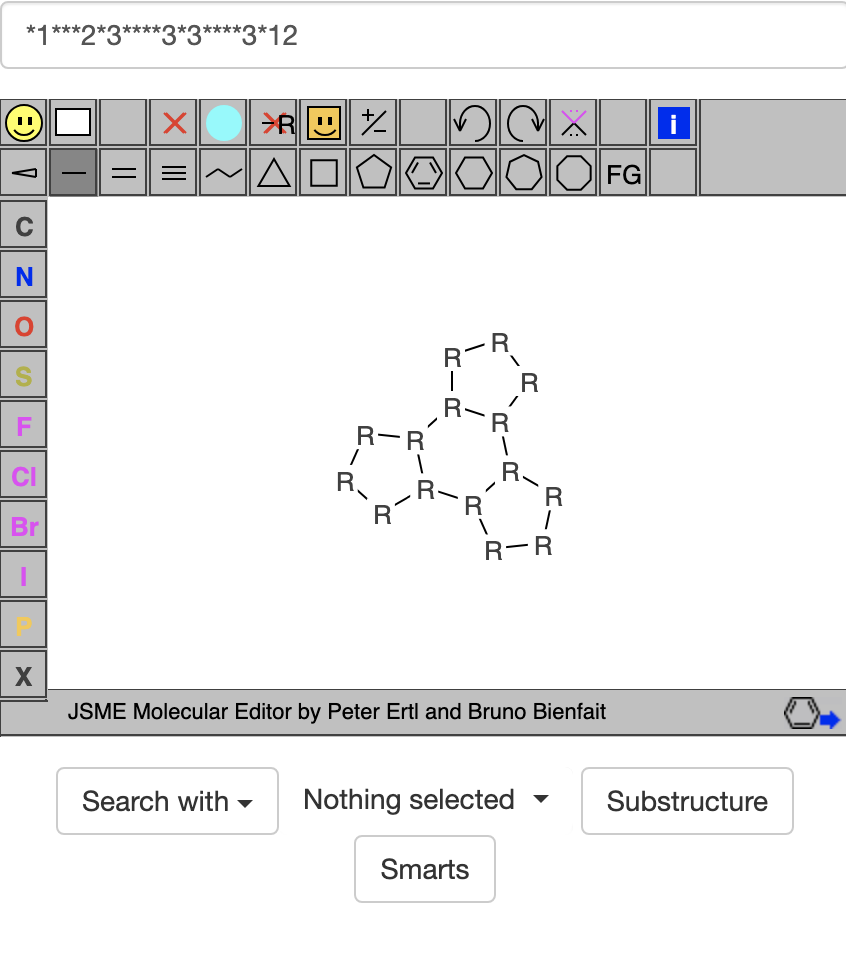
Then I clicked on the SMARTS button below the image to get https://zinc20.docking.org/substances/?sub_id-matches-sma=*1***2*3****3*3****3*12 which gave at least 100 hits, each containing a hexagon surrounded by 3 pentagons.
When I tried to narrow the last search to ones with only 7 hydrogen-bond acceptors using https://zinc20.docking.org/substances/?sub_id-matches-sma=*1***2*3****3*3****3*12&hba=7, it gave 13 hits, but I was surprised that none looked like C1=NNC2C3NN=CN3c3[nH]ccc3N12 shown above. I think this is because C1=NNC2C3NN=CN3c3[nH]ccc3N12 is not itself in the ZINC20 database. I think it is just a scaffold that is part of other compounds in the database.
When I put C1=NNC2C3NN=CN3c3[nH]ccc3N12 in https://zinc20.docking.org/substances/home/ and then pressed the Substructure button under the image box, it gave https://zinc20.docking.org/substances/?sub_id-matches=C1%3DNNC2C3NN%3DCN3c3%5BnH%5Dccc3N12 with 12 hits, each of which contains the 1 hexagon + 3 pentagon core but also looks quite large and complicated, a bit like a Klingon Bird of Prey ship from Star Trek. Note that in the html address, it has converted = in the SMILES code into %3D, [ into %5B, and ] into %5D.
One more thing I tried was https://zinc20.docking.org/substances/?num_rings=4&hba=7&hbd=3, which gave many hits. Some, like ZINC #'s 81470, 90124, 91131, 91284, and 125822, have a cyano group near an -NH2 group, as on the molecule I wanted to approximate at the start of this thread. Comparing this search with some other searches above shows that sometimes ZINC20 uses num_rings=4 to find things with 4 rings, and sometimes it uses ringcount=4 to find them.
I hope this helps!
(edited to remove misleading lines)
Maybe I can also share another strategy on how to put together different substructures here. I usually made myself limit the variety of designs, so I'm not posting the exact designs here.
Using ZINC to explore new scaffolds and combining them together can be useful.
In the Y4R series applying the same logic 80-90% are exact matches to library compounds.
As I open a new puzzle, I inspect the hbond in starting ligand, and try to keep that hbond in any later designs.
Then I'd remove "excessive" parts in the compound, like unsatisfied h-doners and acceptors, halogens etc. Then reconstruct the compound starting from the h-bonding substructure I want to keep. To increase the chance of getting library hits, try to think like synthetic chemist when they need to make a compound. For example, expanding the compound by connecting ring systems with linkers, and probe other h-bonding positions at the same time.
There are many types of linkers, big and small, polar and apolar. But I'm only comfortable to use two kinds of small, polar, rigid linkers (which limits the diversity of my designs). This time the linker positioned in the right h-bonding site, so the exact hits/highly similar hits usually score decent.
I'm not saying using the ZINC20 site is necessary. If you can do well on Ligand Design Puzzles with only the tools inside of Foldit, more power to you. I don't use the ZINC20 site so much myself, but it helps when I am low on ideas. It also seems that some people make much better scoring designs when they don't worry about the compound library and its restrictions. For these people (and for the scientists who want to use their results), it would be good to have ways to convert their non-library designs into on-library ones, and I think the ZINC20 site can help with this.
With that said, some SMARTS code that I find helpful is that you can use [#7,#8] to stand for any acceptor and [#7,#8;!H0] to stand for any donor. For example, in my goal compound N#Cc3c(N)cc(O)c4c1C=NNCc1c2CC(=O)N=Cc2c34, one ring has =N-NH- in it. There the =N- acts as an acceptor while the -NH- acts as a donor. This means I can use [#7,#8][#7,#8;!H0] in place of them to find other acceptor-donor combinations (like -O-NH-) that might work just as well. Another ring has =N-C=O in it. There both the =N- and =O act as acceptors, so I can use [#7,#8]*[#7,#8] in place of the =N-C=O to look for other pairs of acceptors that might work just as well.
If I go to https://zinc20.docking.org/substances/home/ and put [#7,#8]*[#7,#8] in the box above the molecule viewer, I get:
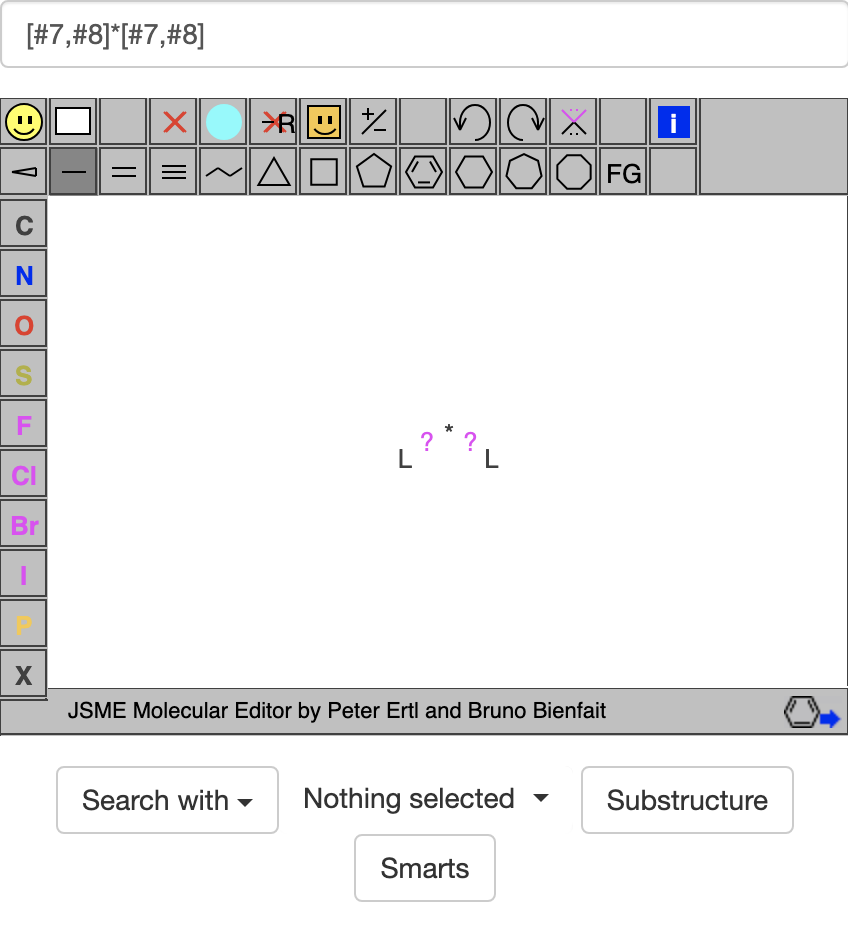
In the image box, it seems the ?'s represent general bonds (could be single, double, aromatic, etc.), the *'s represent general atoms, and the L's stand for general acceptor atoms. If I then click on the SMARTS button, it gives: https://zinc20.docking.org/substances/?sub_id-matches-sma=%5B%237%2C%238%5D*%5B%237%2C%238%5D,
which lists many compounds each containing a pair of acceptors with one atom between them. Note that when making the html above, the ZINC20 site has converted [ to %5B, # to %23, each comma to %2C, and ] to %5D.
If I go to https://zinc20.docking.org/substances/home/ and put [#7,#8][#7,#8;!H0] in the box above the molecule viewer, I get: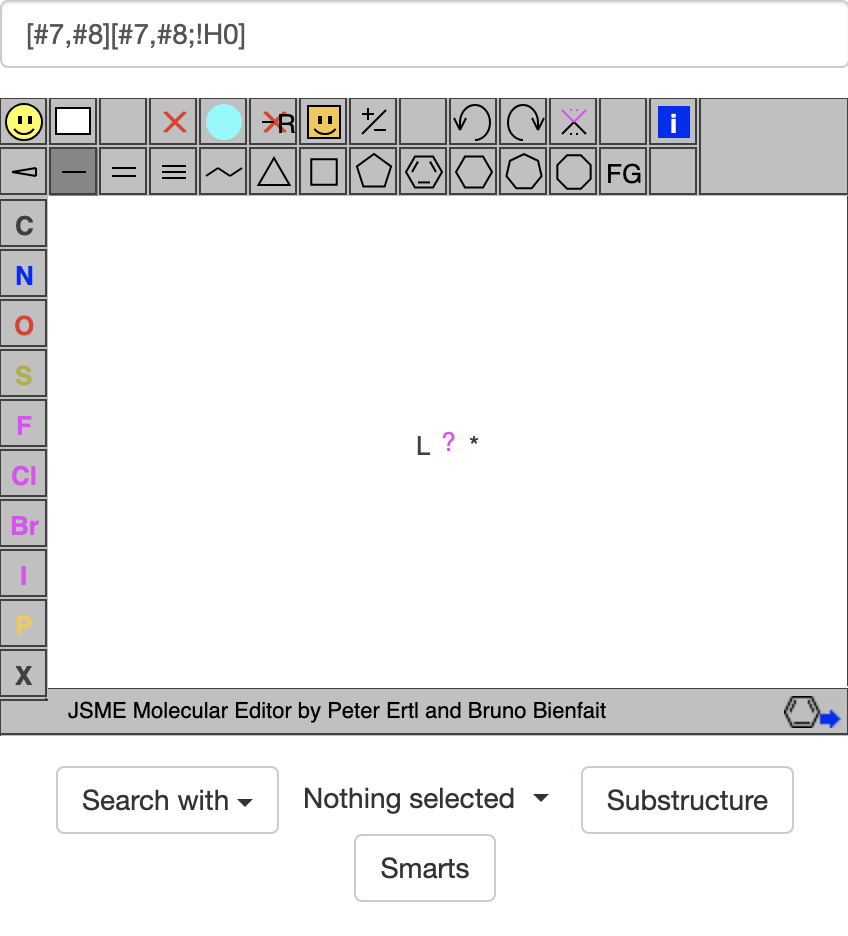
Again, the L stands for a general acceptor atom and the ? stands for a general bond, but here an * stands for the donor atom. If I then click on the SMARTS button, it gives: https://zinc20.docking.org/substances/?sub_id-matches-sma=%5B%237%2C%238%5D%5B%237%2C%238%3B%21H0%5D, which lists many compounds each with an acceptor atom adjacent to a donor atom. Note that when making the html above, the ZINC20 site has converted ; to %3B and ! to %21.
Knowing the html for both [#7,#8]*[#7,#8] and [#7,#8][#7,#8;!H0], I can make a SMARTS search that looks for compounds with both donor-acceptor combinations: https://zinc20.docking.org/substances/?sub_id-matches-sma=%5B%237%2C%238%5D*%5B%237%2C%238%5D&sub_id-matches-sma=%5B%237%2C%238%5D%5B%237%2C%238%3B%21H0%5D. This particular search gives only 18 hits, and the ones with ZINC #'s 5904950, 44930779, and 334138525 might be reasonable replacements for about half of my goal compound.
Earlier I discussed a search using https://zinc20.docking.org/rings/?ringcount=4&hba=7&hbd=3 to look for compounds with 4 attached rings, 7 acceptor atoms, and 3 donor atoms. When I did this search, I didn't realize that the rings part of the ZINC20 site was looking for patterns instead of compounds. This means it only considers things that are part of the ring system, not attached side groups like my goal compound's -OH, -NH2, and cyano groups. So for searching the rings part of the ZINC20 site, I only need to consider 4 of the acceptor atoms and 1 donor atom. Using the search https://zinc20.docking.org/rings/?ringcount=4&hba=4&hbd=1 gives many interesting hits, like c1cnc2c(c1)c1nc[nH]c1c1cccnc12 below: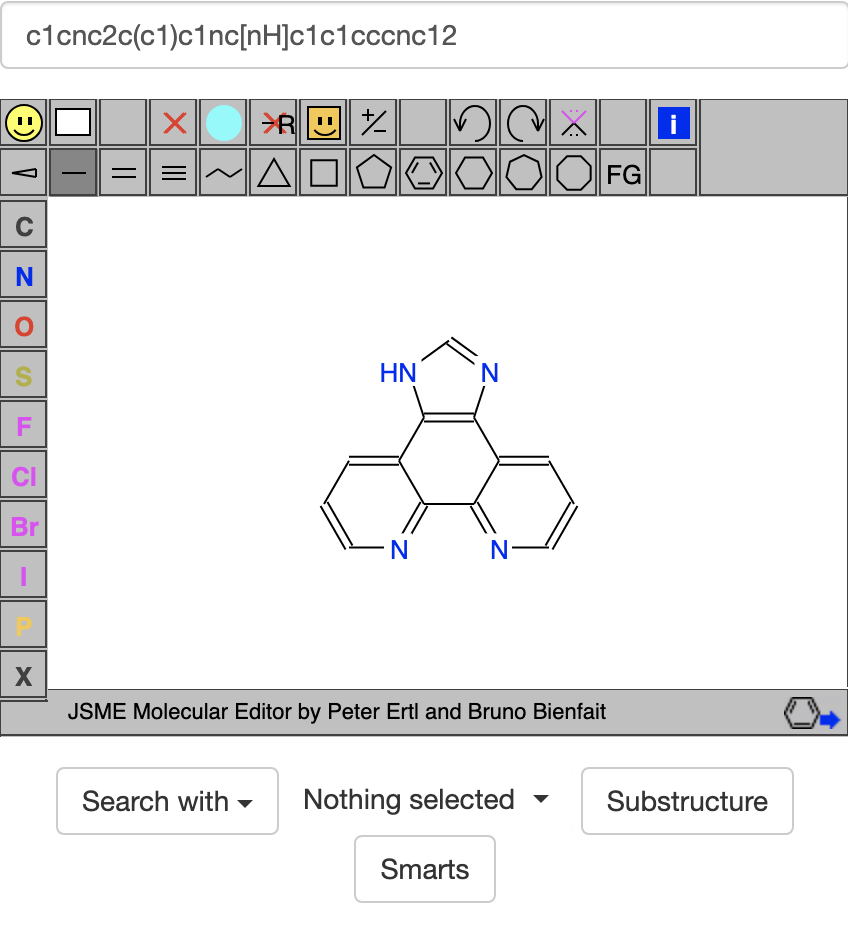 To see even more results, change the html to https://zinc20.docking.org/rings/?ringcount=4&hba=4&hbd=1&page=2 or https://zinc20.docking.org/rings/?ringcount=4&hba=4&hbd=1&page=3. I don't know how many pages there are, but p.3's
To see even more results, change the html to https://zinc20.docking.org/rings/?ringcount=4&hba=4&hbd=1&page=2 or https://zinc20.docking.org/rings/?ringcount=4&hba=4&hbd=1&page=3. I don't know how many pages there are, but p.3's c1ccc2c(c1)NC1(CCCCC1)n1ncnc1-2, O=C1OCCC2C1C1CCCN1C21CCNCC1, and O=C1c2ccccc2-n2cncc2C2CCNC12 look interesting to me, and p.5's O=c1[nH]nc2c3ccccc3n3c(c1-2)CCC3 does too.
If you are curious about systems with multiple rings that are not all fuzed together as in the above searches, try searches like https://zinc20.docking.org/scaffolds/?ringcount=4 or https://zinc20.docking.org/scaffolds/?ringcount=4&page=2 instead.







 To see even more results, change the html to
To see even more results, change the html to