A huge thank-you to everyone who participated in our ten-round Drugit VHL series (20 Oct 2021 – 12 Jan 2022). Across those puzzles, 333 of you submitted about 6,500 unique molecules. The Meiler lab team, along with our collaborators at Boehringer Ingelheim selected 19 of your designs for synthesis—and one of them went on to bind the VHL E3 ligase as predicted in-game, a result now confirmed by X-ray crystallography.
This milestone appears in Nature Communications under the title “Drugit: crowd-sourcing molecular design of non-peptidic VHL binders.” You can read the open-access paper here: https://www.nature.com/articles/s41467-025-58406-0 and check out the Drug Design Minute here: https://youtube.com/shorts/I0xn6vckcfM
Your creativity shows that crowdsourced intuition can push drug discovery forward!
Congratulations to the authors, their institutions and all Foldit users who contributed.
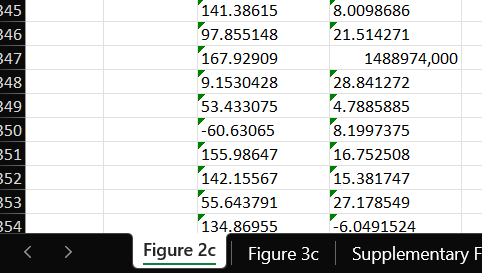
I find there is inconsistent (incorrect ?) data in the supplementary info spreadsheet, tab 'Figure 2c', multiple cells show large numbers whereas other cells show fractional values with US decimals. Can you confirm that the values used in the paper are within a proper range (<1000) ? I did not edit these values when loading your spreadsheet. If the value had been 14.88974 why did it not load similar to the other values around it ? Has this been checked by the reviewers ?
May I also suggest not saving excel sheets as a macro-enabled sheet but instead as a regular excel 97 compatible sheet to avoid unintentional macro-virus spreading and maintain compatibility with non-Microsoft spreadsheet products ?
See:
It looks like Contenders lead this series of puzzles. Congratulation for the group that pioneered this kind of puzzles.
Congrats to folder @Nicm25 for his compound that showed binding!!!!!
i noticed a comment in the paper. "molecules with any atom more than 10 Å away from any starting molecule binding site atom were removed." can we get more clarity and more visibility in future puzzles on what this looks like? I don't really want to waste a lot of time, electricity, and computing time trying possible solutions if they would just get thrown out.
maybe i missed it but i swear the "binding either to Ser111 or His115" is the first time i heard there was any binding site or requirements.
"molecules with any atom more than 10 Å away from any starting molecule binding site atom were removed." can we get more clarity and more visibility in future puzzles on what this looks like?
We'll certainly try to be as up-front about any such requirements as we can be.
In this particular case, this filter was added at late stage (after the puzzle series closed), mainly as a way to easily rule out compounds which either weren't touching the protein at all, or which were in the binding pocket and unfeasible massive (and thus were not only poor by the molecular weight objective but also would never be passed through to synthesis later anyway). Relatively few compounds were ruled out by this filtering.
The nature of the target we gave you meant there really was only a single binding pocket where you'd get decent scores (as trying to bind elsewhere meant that trying to bind outside of the pocket would likely mean that the molecular weight objective went down faster than the interaction energy score component went up).
I wanted Thank you all for the amazing research work and allowing us to get this credit; our names side by side with some of the greatest minds around the world in this field. Respect. Kia Kaha ` Aotearoa





