In the current series of CACHE puzzle, the SMILES string is now revealed for each compound in the ligand queue.
Many external tools accept the SMILES string. Having the SMILES string is useful for viewing and searching.
The current ligand queue requires manually transcribing the SMILES string displayed. This is time-consuming and error-prone.
The SMILES string is currently available only for the ligand queue. The SMILES for the current solution is not shown. The SMILES for compound library matches are also not shown.
The SMILES string should be available everywhere.
For the compound library, the SMILES string should be shown with the properties of the submitted compound in the Compound Library window. The SMILES should also be available for the results in the Load Library Compounds window. It should be possible to copy the SMILES string from each of these locations.
Outside of the compound library functions, the SMILES string should be shown in the Segment Information window for the main window compound. The Segment Information window already has a text box for segment notes. A second text box could be added for the SMILES string, or there could be an option to set the segment note to the SMILES string.
Saved solutions could also reveal SMILES, but just loading the solution to look at the SMILES would be an acceptable alternative.
It is possible to glean SMILES information from log.txt, but this usually requires shutting down Foldit. The SMILES in log.txt use the long format, with all hydrogens explicitly included, making it difficult to determine which SMILES is which.
There have already been multiple requests for Foldit to accept SMILES as input on puzzles which involve designing small molecules.
On Windows machines, it is possible to view the contents of an *.ir_solution file using Notepad. Some of the text is human-readable and gives useful information. Why not include a human-readable line somewhere in each *.ir_solution file for its ligand's SMILES code?
A LUA command like structure.GetSMILES that returns a string containing the SMILES code for the ligand would be helpful too.
I agree 100% with this post
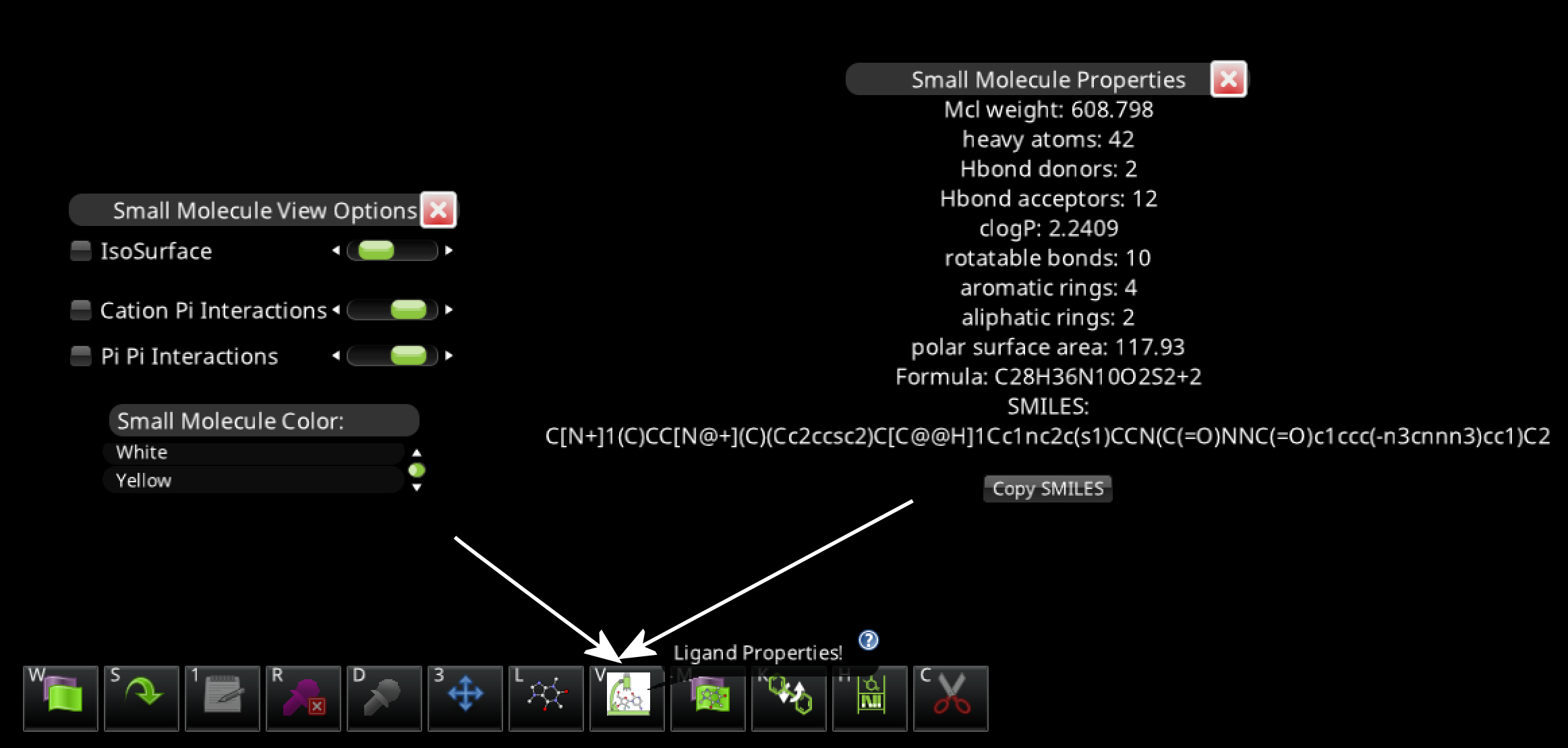
The latest devprev, V46-20250213-cae7350b02-win_x64-devprev, displays the SMILES string in the Small Molecule Properties window.
There's also a "Copy SMILES" button, which copies the SMILES string to the clipboard.
There's still no way to input a SMILES string, but this is progress.
The SMILES string will usually be too large to fit in the Small Molecule Properties window, but dragging the window over a dark background makes it visible.
The Small Molecule Properties window requires selecting the ligand, then using the microscope-and-molecule icon or the "v" hotkey. This opens both the Small Molecule View Options and Small Molecule properties windows.
Since it is relatively easy (but time consuming) "visual copying" small molecules, and since the Library implicitly "copy-paste" a compound to Foldit, it makes sense (for players to save hand fold time) to get a tool to import SMILES.
This would speed up te process of trying (external sources) compounds, build on an existing molecule (SMILE or library imported) to find the good binding one, using external tools to compare or create new candidates etc.
My dream would be to make a recipe that would automatically try a lot of "player's intuition" selected compound imported with their SMILE format.
This doesn't prevent a week of Foldit work to calculate the best position of the pre-selected ones.




