LociOiling Lv 1
Back to the question of how the compound library works….
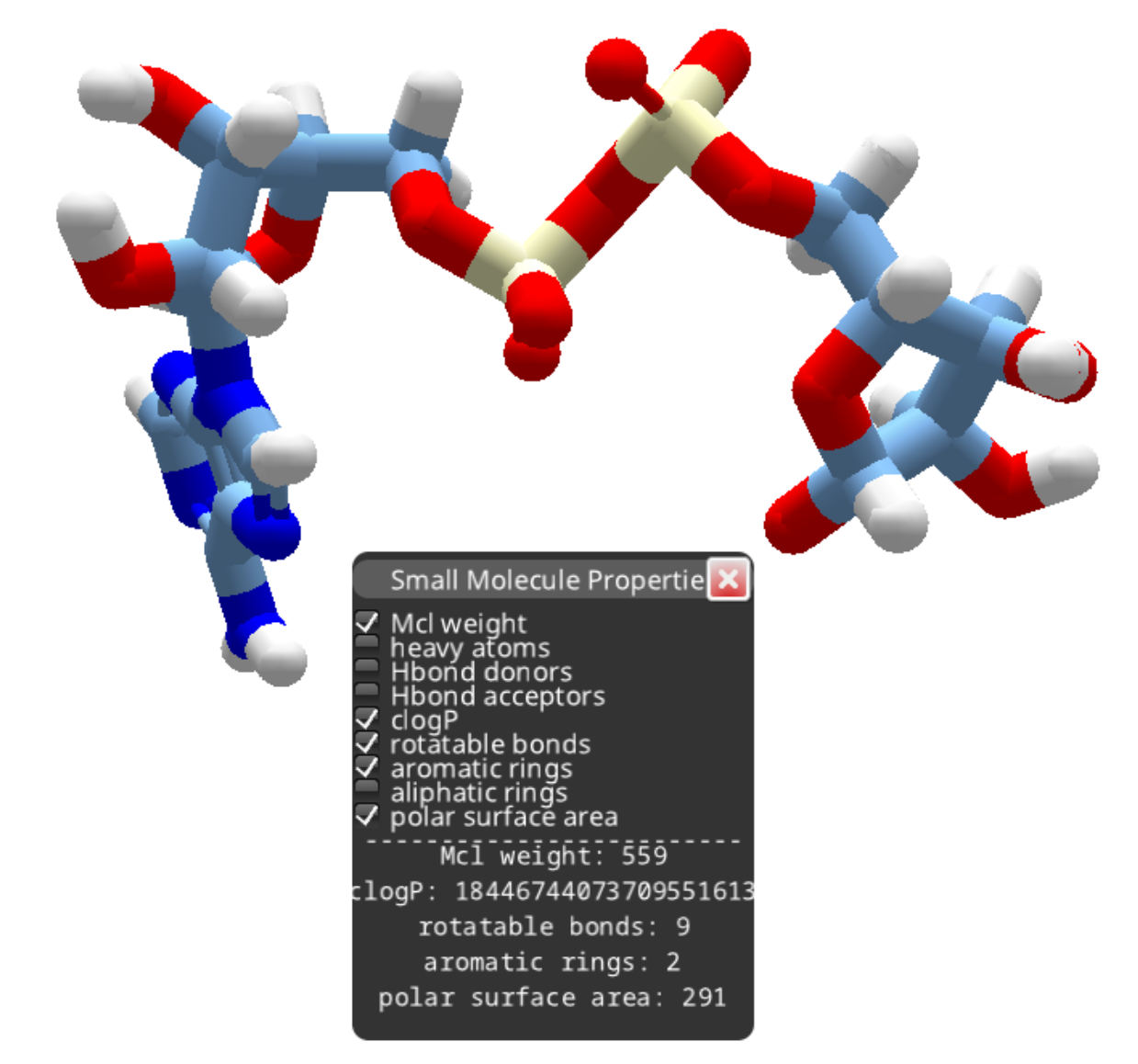
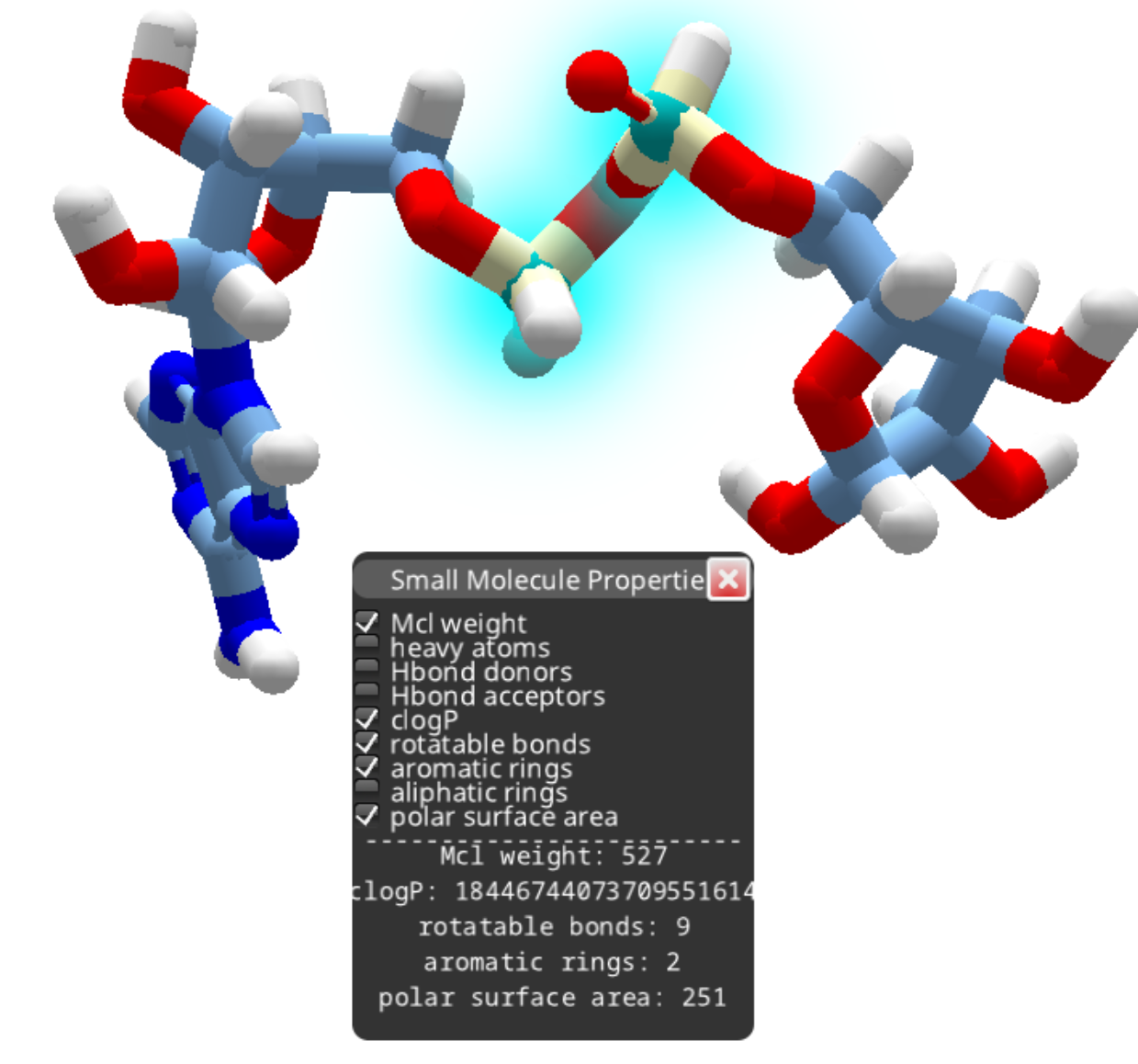
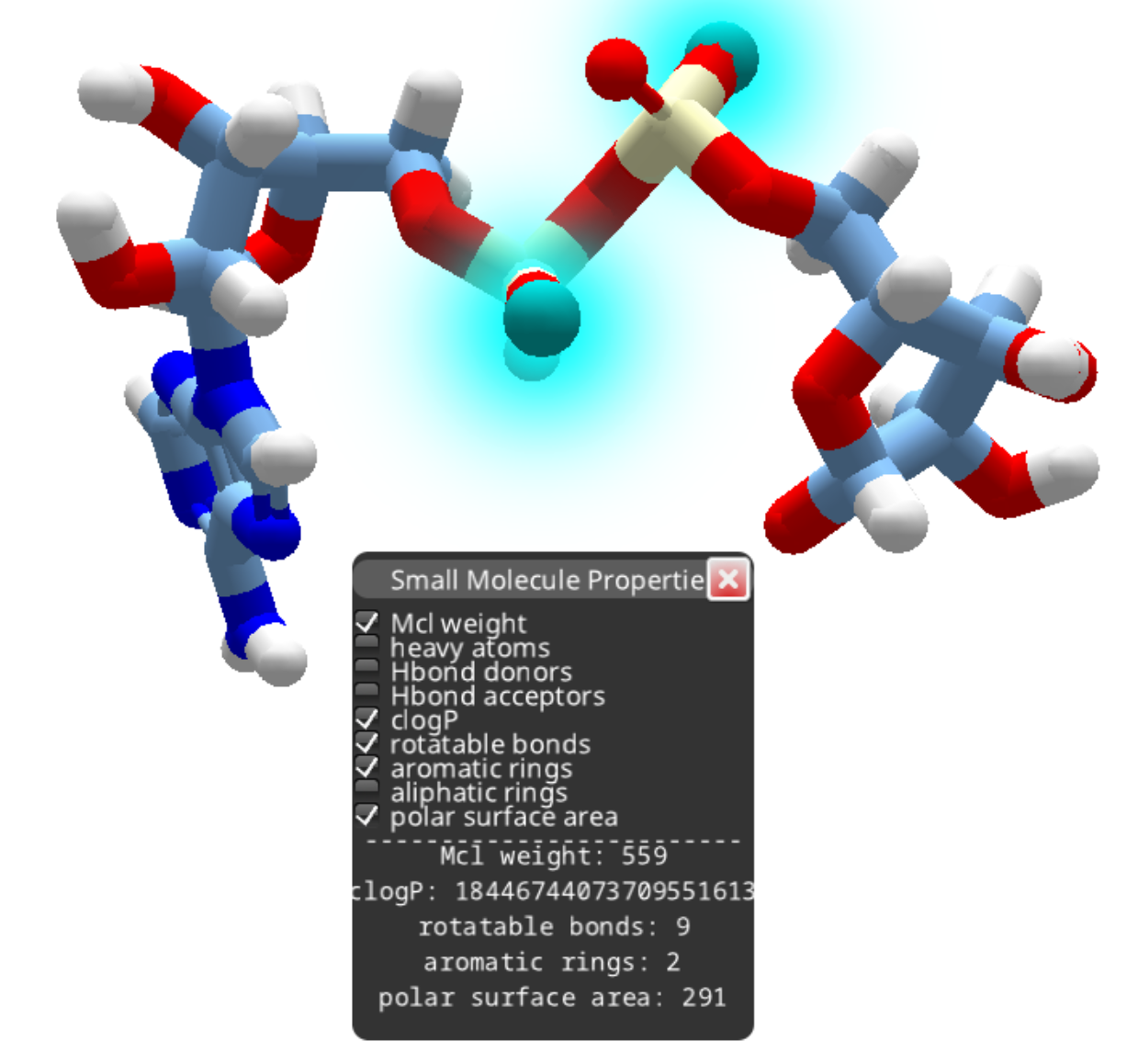
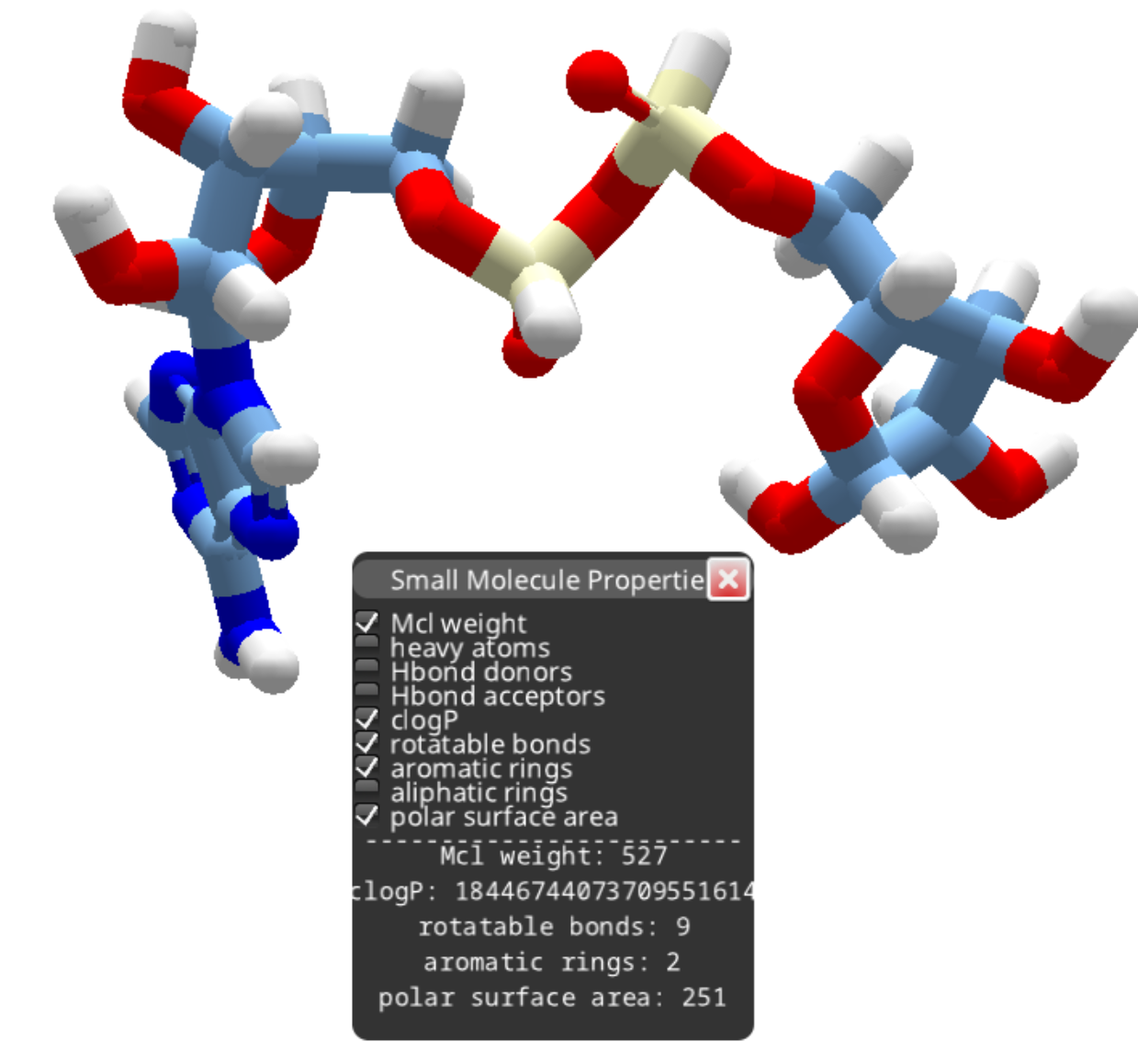
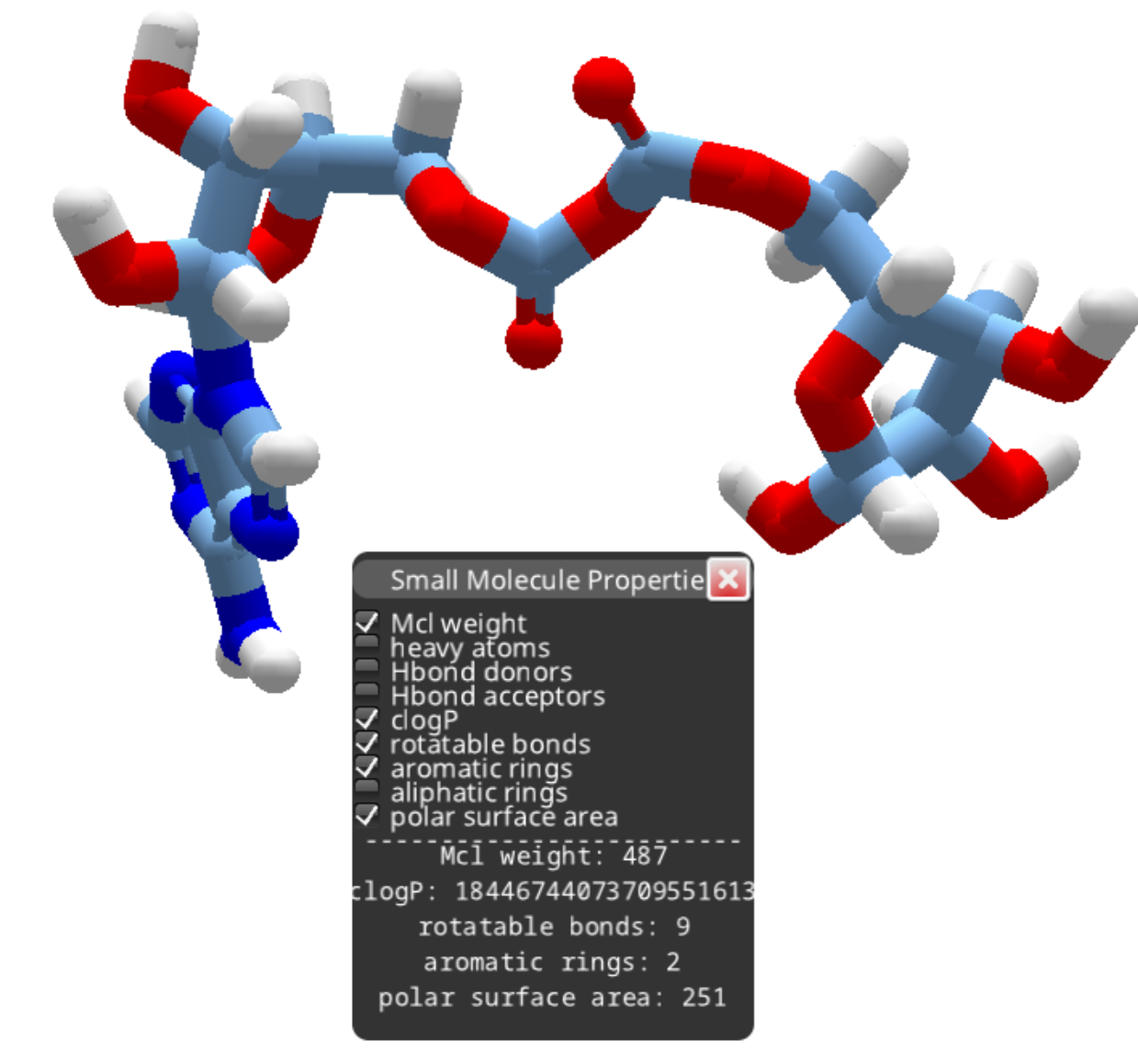
After removing the phosphorous as outlined above, you can continue deleting atoms to produce a smaller compound.
With the ligand selected, you can then open the Compound Library panel (hotkey "H") and use "Upload Current" to start a search. You'll need to give your new molecule a name.
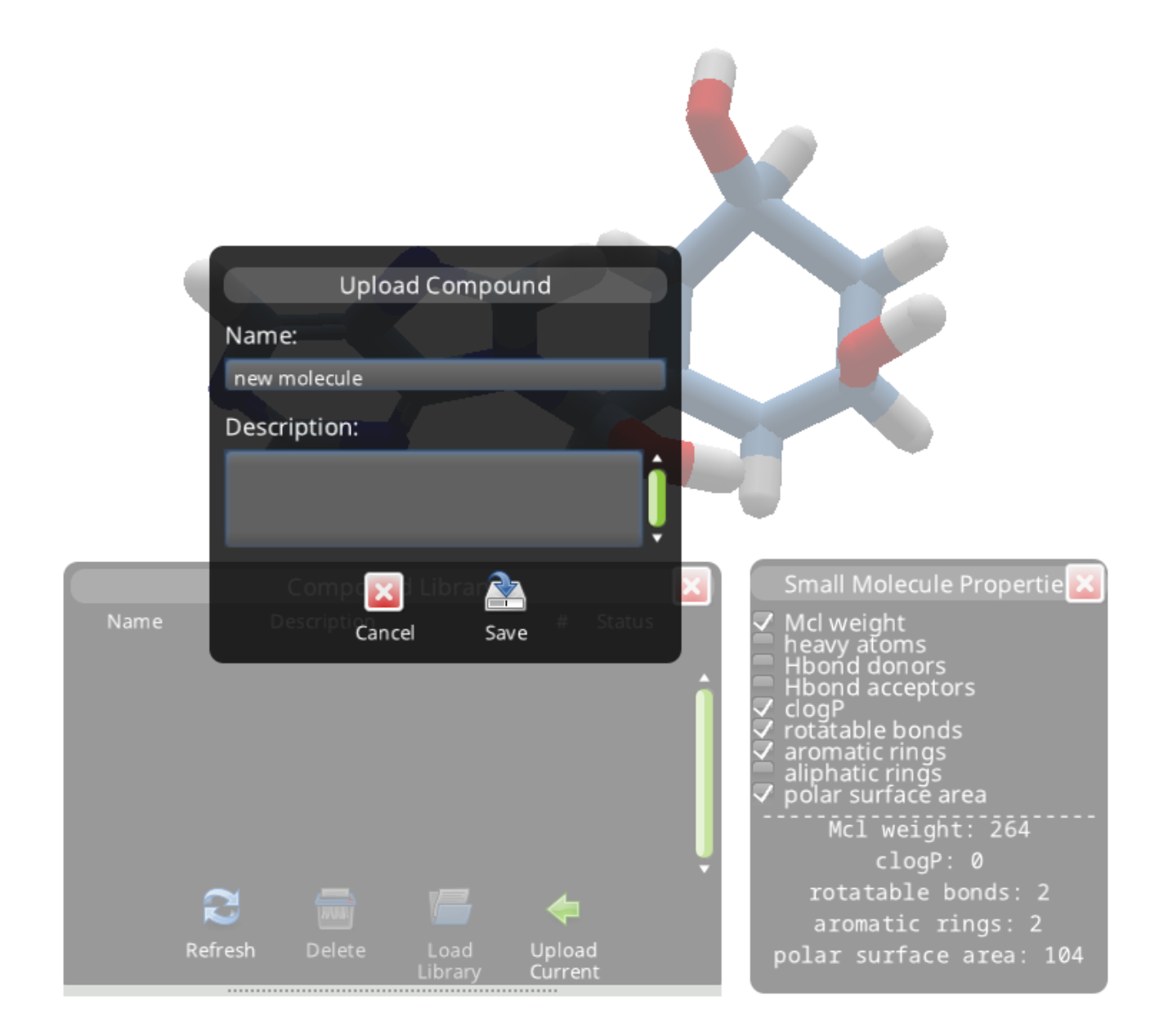
The panel shows your submission is "Running".
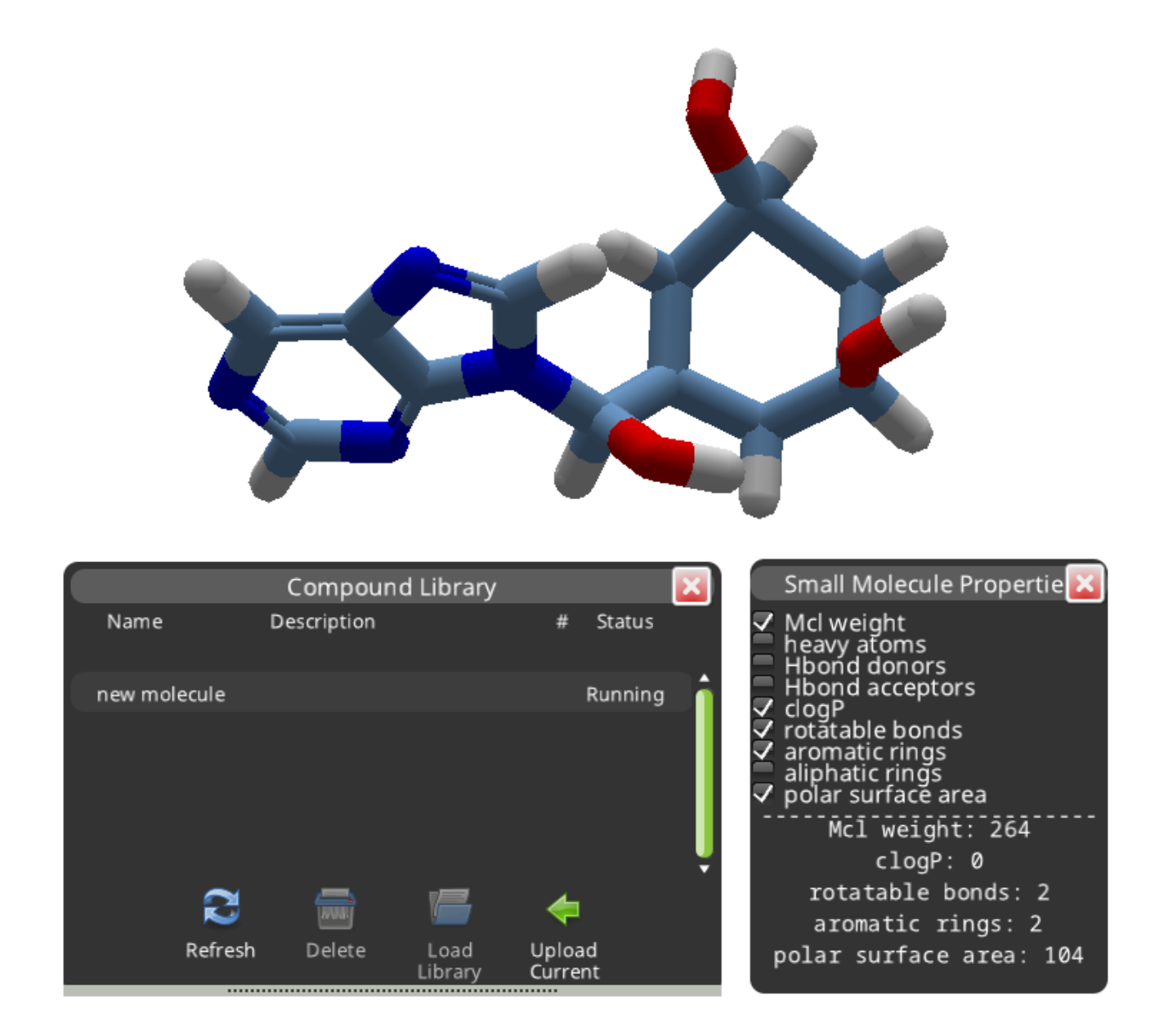
A search normally takes a few minutes before it's "Done". The number of results is shown in the "#" column. This search found 35 results.
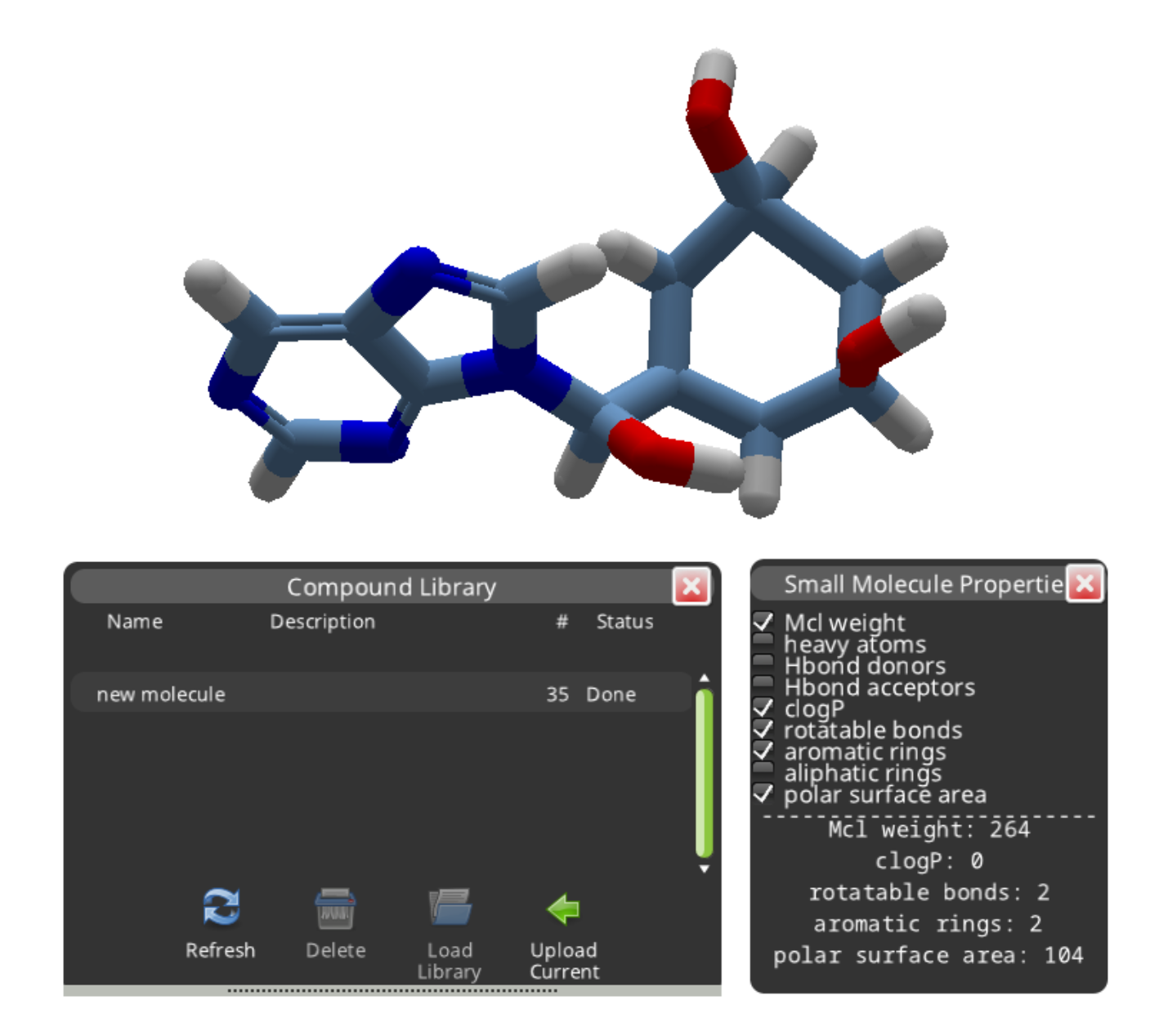
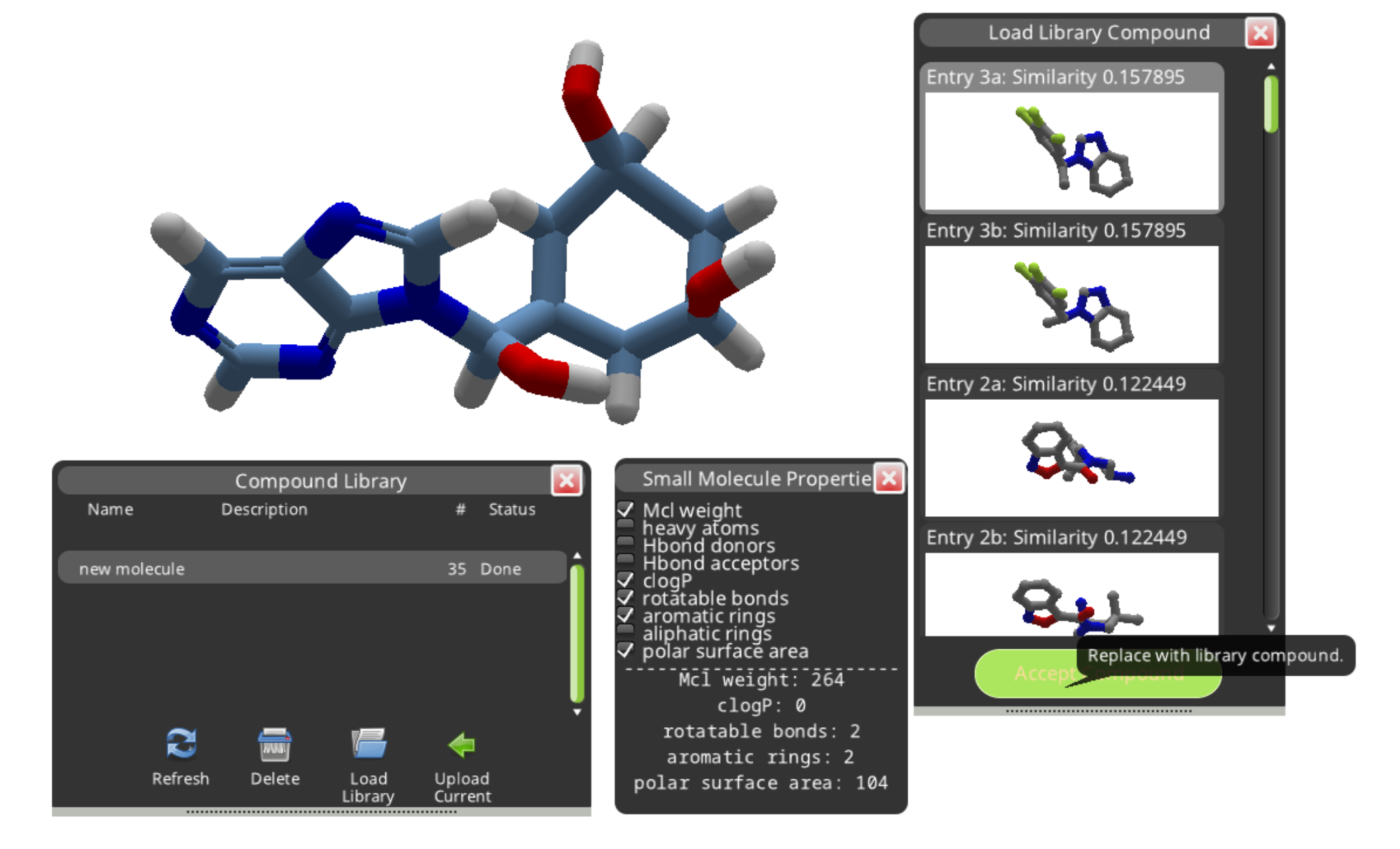
The "Load Library" button displays the results in a separate panel. The results are shown in order of decreasing similarity to the compound you submitted. Selecting a result and clicking "Accept Compound" replaces your new molecule with a library compound.
It's a good idea to create a segment note to identify the new compound.
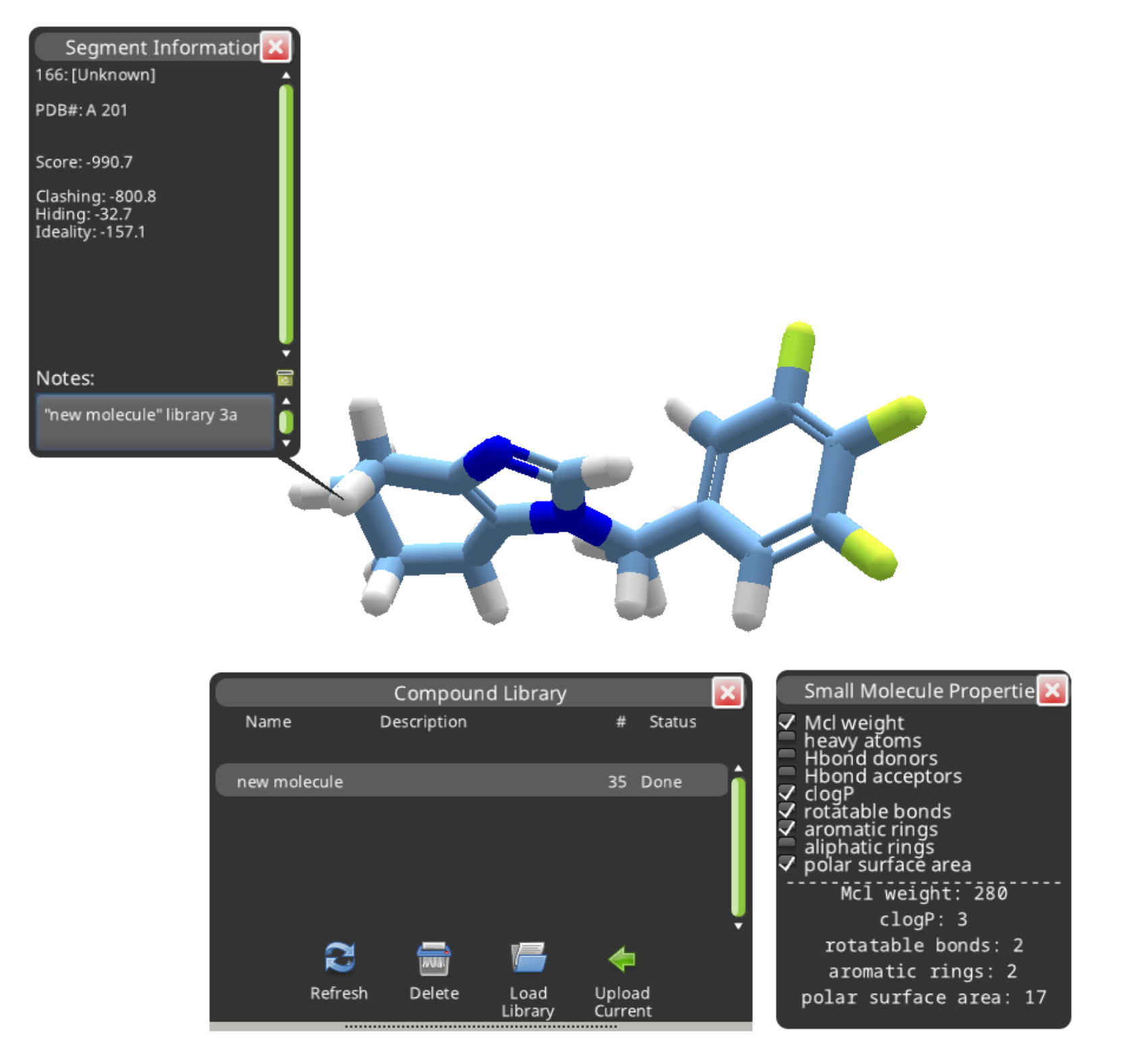
Segment notes can be hidden or displayed using the hotkey "1".
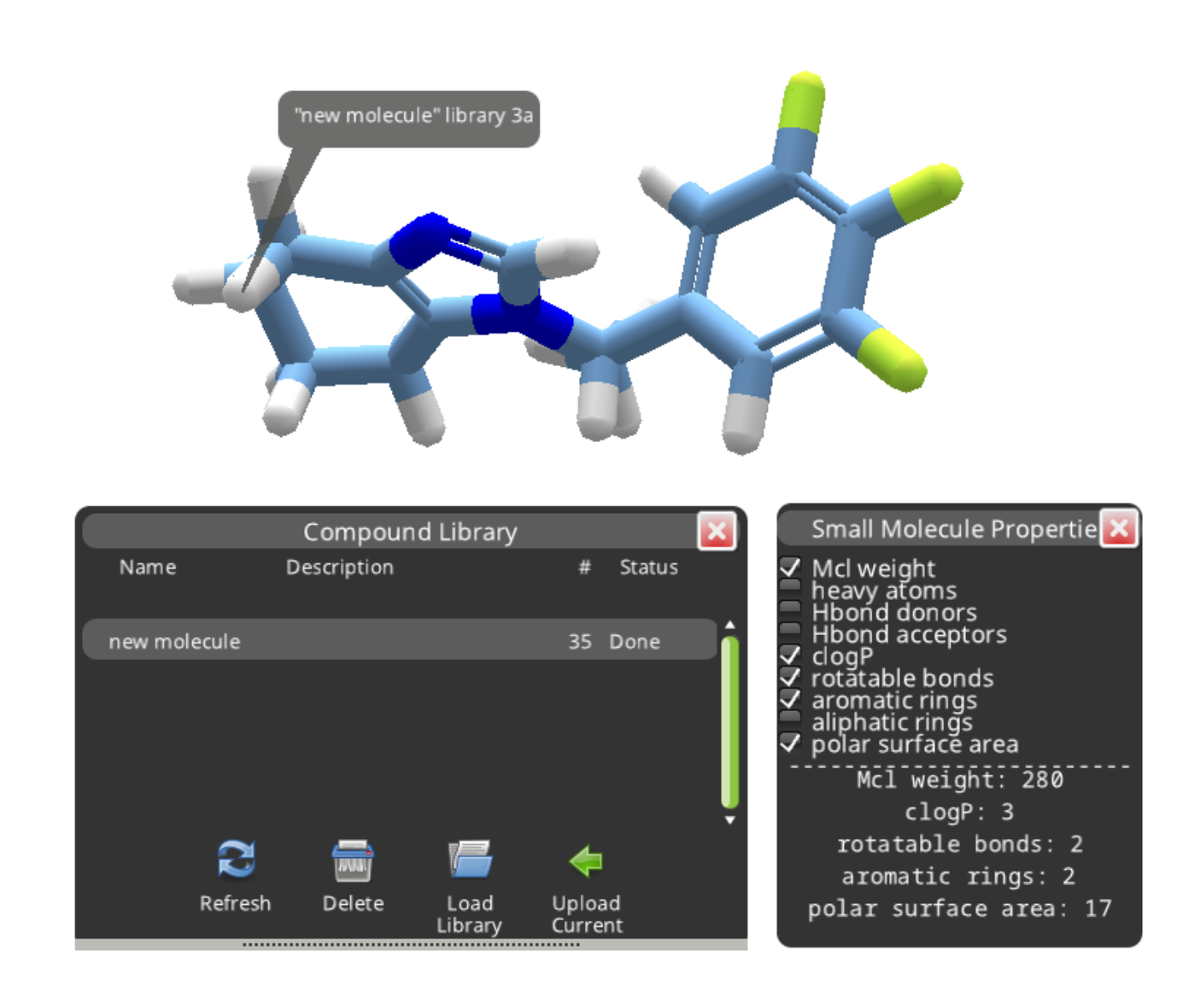
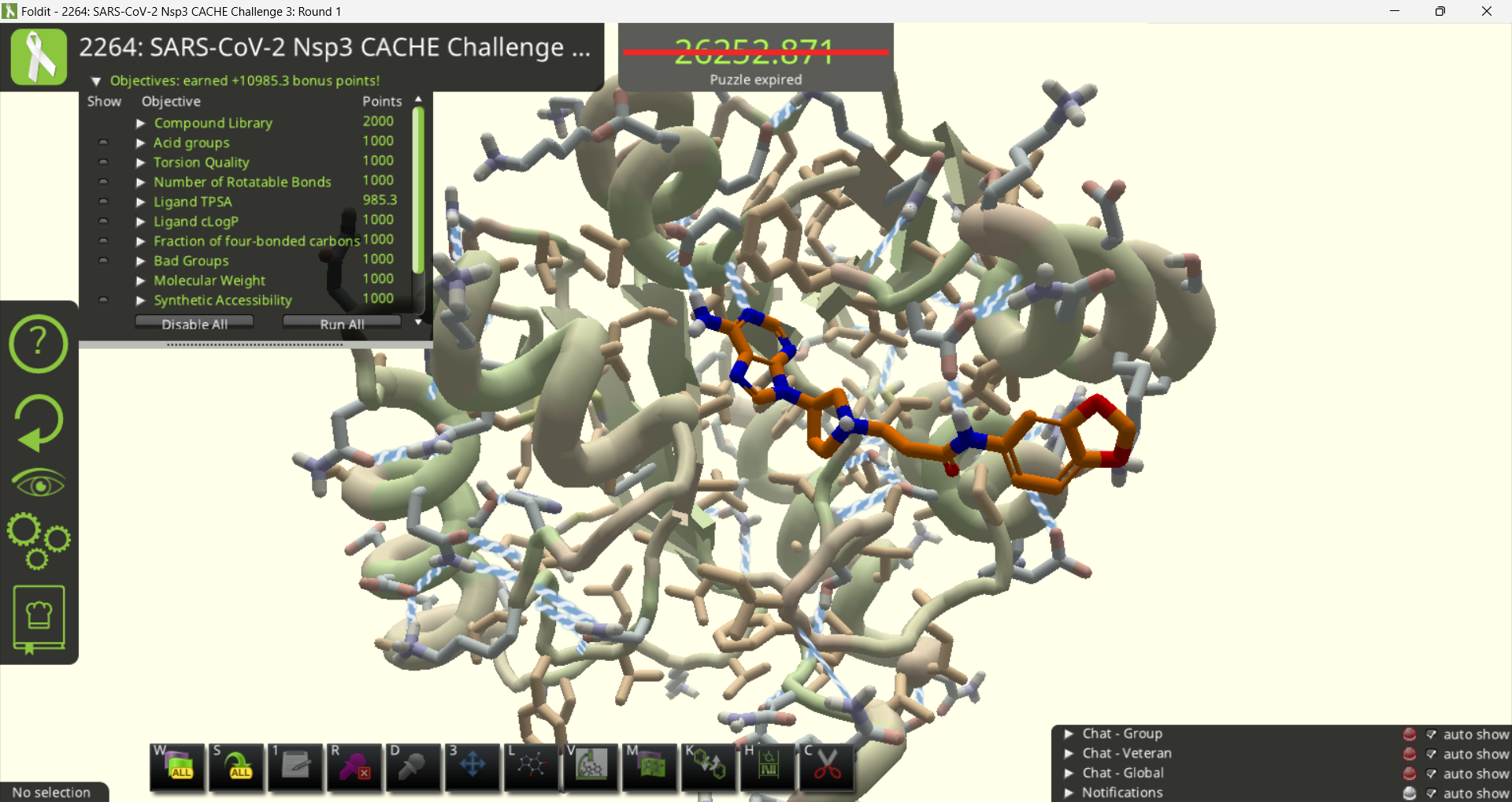
It's also a good idea to take a look at the Objectives. Compound 3a was only a weak match (0.15) for the submitted molecule. It incurs a big "Bad Groups" penalty, and the cLogP is also a little high.
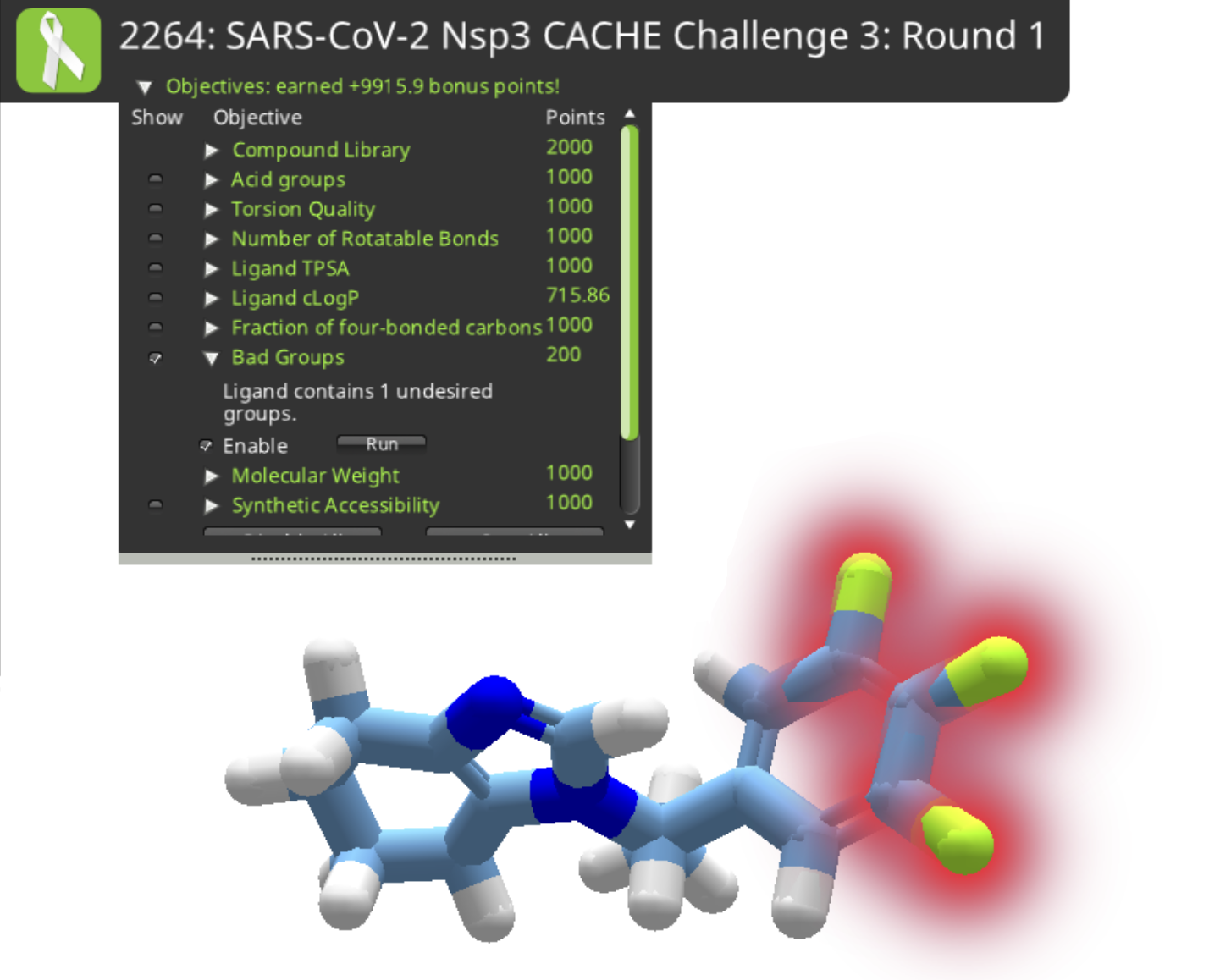

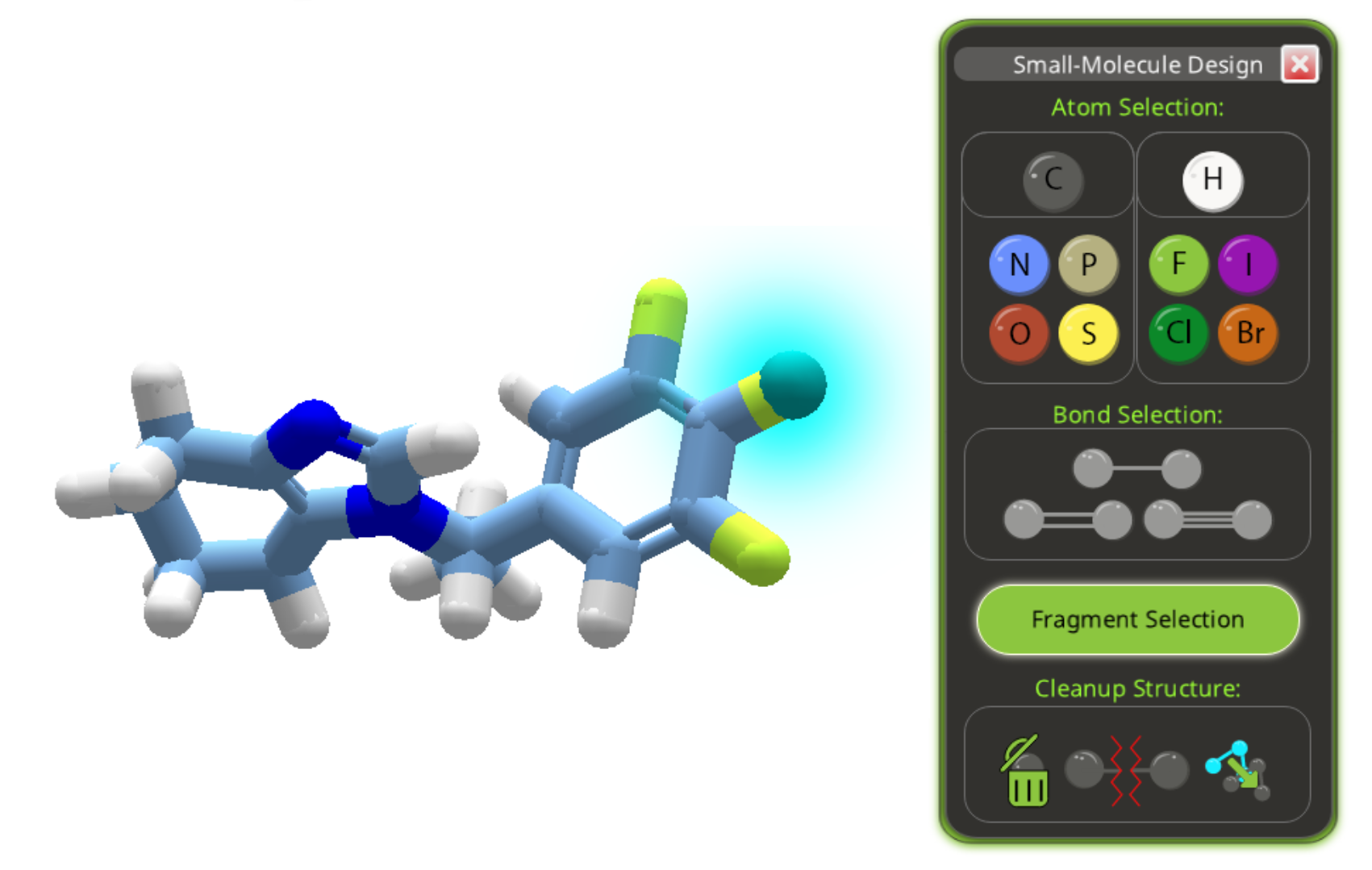
The three yellow atoms on the right of compound 3c are highlighted as a bad group. There's no way to directly identify which element they are. They look like sulfur, but a sulfur would normally have a hydrogen attached. The solution is to open the ligand design tool, select one of the atoms, and try substitutions. A successful substitution will have the same molecular weight and of course the same color as the mystery atom. It turns out the three mystery atoms are fluorine in this case. While some drugs do in fact contain three fluorine atoms, any fluorine is going to earn a "bad groups" deduction for this puzzle.