
rmoretti Staff Lv 1
Objectives
Maximum bonus: +1 000
Torsion Quality (max +1000)
Keeps bond rotations in a good range. Using Wiggle or Tweak Ligand can fix bad torsions. (Show highlights torsions to be rotated.)

Closed since almost 2 years ago
Intermediate Intermediate Intermediate Intermediate Intermediate Intermediate Intermediate Intermediate Intermediate Overall Overall Overall Overall Overall Overall Overall Overall Overall Small Molecule Design Small Molecule Design Small Molecule Design Small Molecule Design Small Molecule Design Small Molecule Design Small Molecule Design Small Molecule Design Small Molecule DesignNew CASP ligand target just dropped!
This puzzle is part of the CASP16 competition. Foldit players are participating to see how well they can predict how small molecules can bind to proteins. Note that in contrast to prior drug design puzzles, we're not looking to design a new molecule, but instead are interested in getting a good structure for the provided ligand compounds.
The protein target is the WDR55 domain of Human LRRK2. This is the same protein that was the target for the CACHE Challenge #1 ... in fact, the ligand in question is one of the CACHE participant designs. (Though as this was a CACHE Challenge Foldit did not participate in, it isn't one of your designs.) There's only 1 ligand of interest in this competition, but the CASP organizers provided a "racemic" specification -- this means that the actual ligand structure in the crystal is one of two possibilities. The starting small molecule is one of the two possibilities, and is placed somewhat arbitrarily in the middle of the protein. You can use the Ligand Queue tool to get the other possibility, which differs in which way a methyl group comes off the molecule.
Note that we don't have a good idea of where the binding site of this protein is -- one of the reasons this was a good target for the inaugural CACHE Challenge is that the protein didn't really have any known ligands. As such, you'll need to be thorough about searching for the best binding location for this ligand.
As before, we've allowed full backbone and sidechain flexibility on this puzzle, but the since the backbone is very unlikely to change at all from the starting conformation there a restraints (unchangeable bands) to the starting conformation -- these will show up as red lines if you move the backbone too far.


Maximum bonus: +1 000
Torsion Quality (max +1000)
Keeps bond rotations in a good range. Using Wiggle or Tweak Ligand can fix bad torsions. (Show highlights torsions to be rotated.)

I thought CACHE was supposed to get protein-ligand complex crystal structures for the best ligands participants designed.
Did they not get the crystal structure for this ligand bound to the protein?
https://cache-challenge.org/results-cache-challenge-1 says:
"The preliminary crystal structure (refinement in progress) of one compound in complex with LRRK2 shows the compound occupying an unexpected side-pocket, which may be a crystallographic artifact, as binding is still observed in solution when a nearby residue is mutated"
and cites https://cache-challenge.org/sites/default/files/downloadable/forms/crystal_structure.pdf
and https://cache-challenge.org/sites/default/files/downloadable/forms/Preliminary_LRRK2_CACHE_1193_26.pdb
The pdf file above has a picture of the ligand CACHE_1193_26 (with 1F 3N and 2O atoms), so it should be possible to get its SMILES code and compare it with XS381774a CC1Cc2cc(ccc2O1)c1ccnc2ncnn12 (with 0F 4N and 1O atoms) as given at https://predictioncenter.org/casp16/target.cgi?id=361&view=ligand (already I can see that the ligands differ).
The pdf file above also shows the ligand bound near the dimer's monomer/monomer interface and residue G2385
instead of near the protein's central cavity.

I just copied the 2 figures below from the Arthor website.
(1) is for CACHE_1193_26 = Z2141481559 = C19H22FN3O2 with Molecular weight 343.394 and comes from https://arthor.docking.org/index.html#real-database-22q1/sim:/s/C12=CC(F)=CC=C1NC=C2C4CCN(C(=O)C3CC(=O)NCC3)CC4
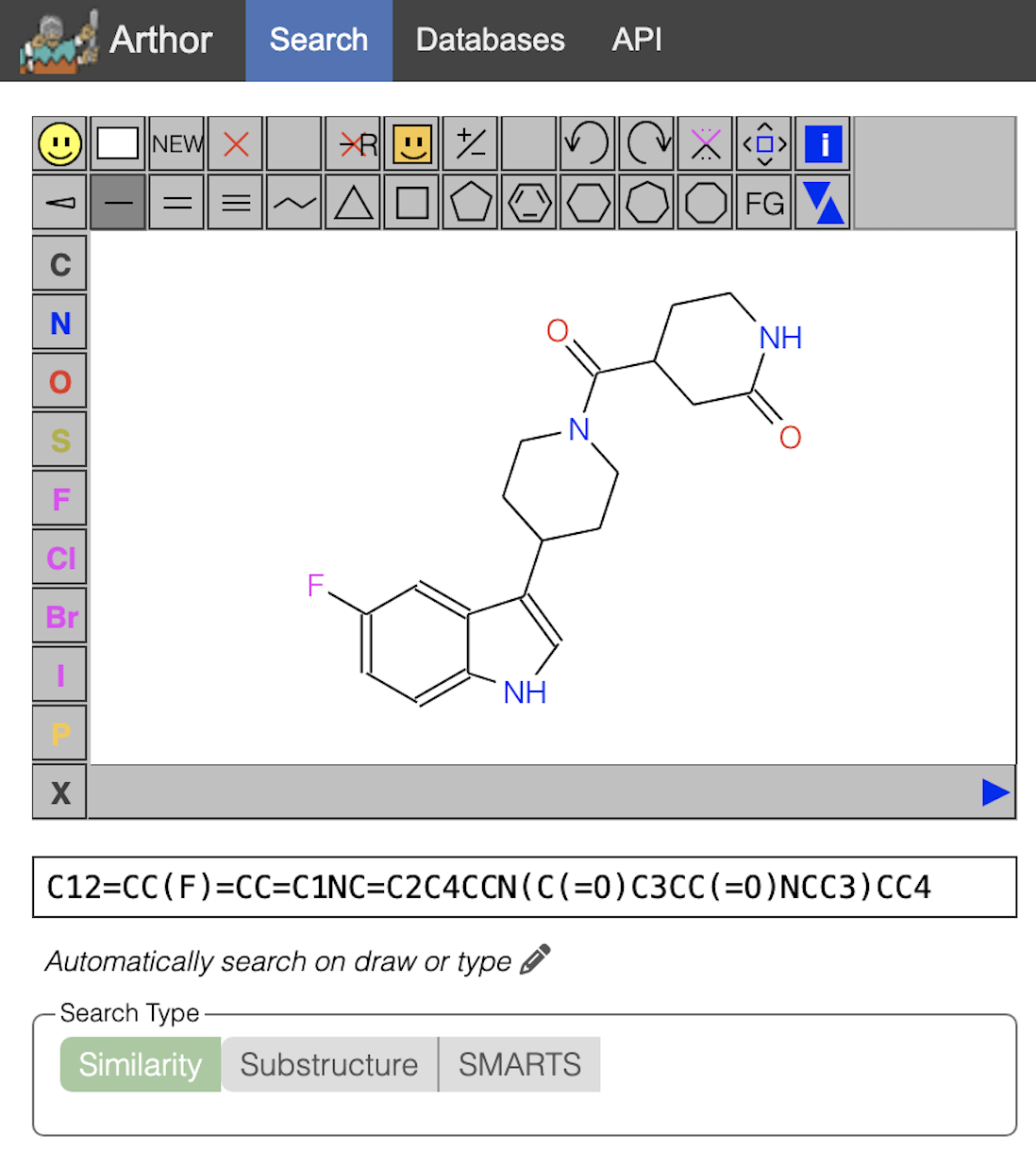
Another SMILES code for the ligand above in (1) is O=C1CC(C(=O)N2CCC(C3=CNC4=CC=C(F)C=C34)CC2)CCN1.
(2) is for XS381774a = CC1Cc2cc(ccc2O1)c1ccnc2ncnn12 = Z2035258982 = C14H12N4O with Molecular Weight 252.271 and comes from https://arthor.docking.org/index.html#real-database-22q1/sim:/s/CC1Cc2cc(ccc2O1)c1ccnc2ncnn12 or https://arthor.docking.org/index.html#real-database-22q1/sim:/s/CC4CC3=CC(C1=CC=NC2=NC=NN12)=CC=C3O4
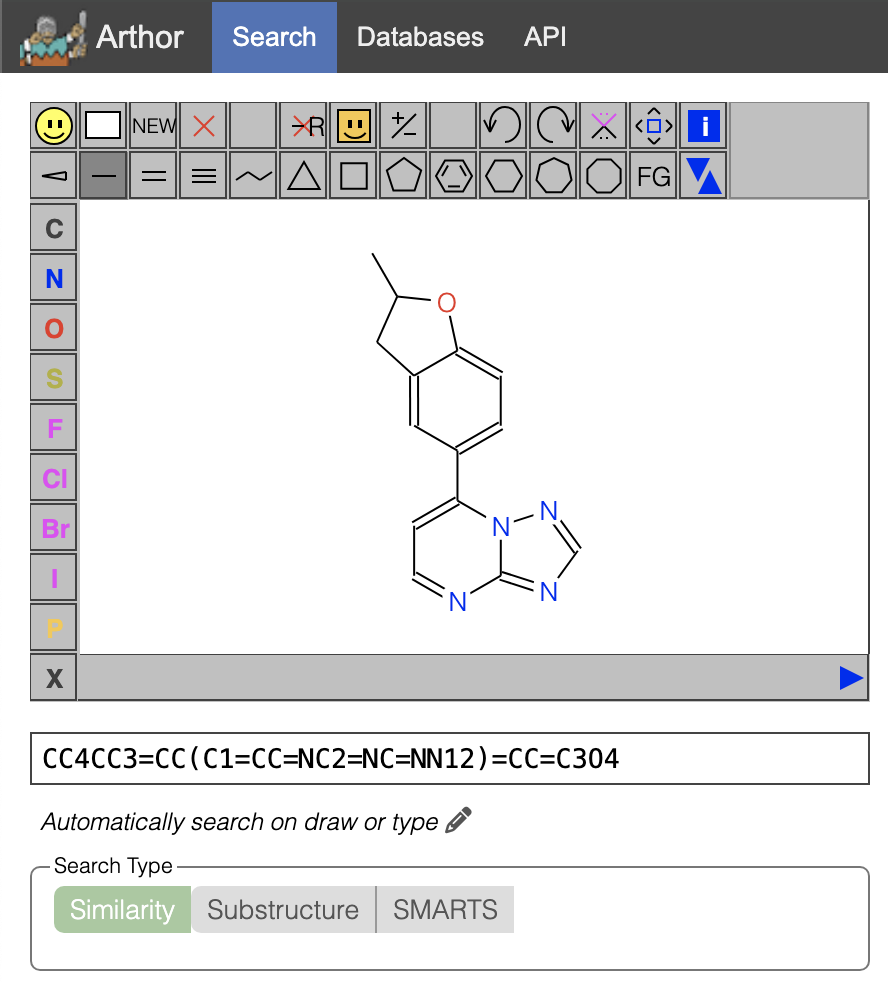
Another SMILES code for the ligand above in (2) is CC1CC2=CC(C3=CC=NC4=NC=NN34)=CC=C2O1.
Sorry if too much information. The Arthor site seems able to read & write multiple SMILES codes for the same ligand.
Some list aromatic C or N atoms with explicit double bonds (=) between them. Others list aromatic C or N atoms as c or n.

https://cache-challenge.org/cache-publications lists several 2024 publications about LRRK2.