949: CASP11 Target Ts818: Sparse Data Contacts
Closed since almost 12 years ago
Intermediate Intermediate Intermediate Intermediate Intermediate Intermediate Intermediate Overall Overall Overall Overall Overall Overall Overall Prediction Prediction Prediction Prediction Prediction Prediction Prediction CASP11 CASP11 CASP11 CASP11 CASP11 CASP11 CASP11 Predicted Contacts Predicted Contacts Predicted Contacts Predicted Contacts Predicted Contacts Predicted Contacts Predicted ContactsSummary
- Created
- July 26, 2014
- Expires
- Max points
- 100
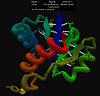
For this target, the CASP11 organizers have provided "sparse" experimental data about possible contacts in the protein. Contacts are scored with the Contact Map filter in this puzzle, and there is an upper limit of 2250 possible points from the filter. The starting structure is an extended chain, and secondary structure predictions can be found below in the puzzle comments.
Top groups
-
100 pts. 11,494
-
-
-
-
-
-
-
-
-
Comments