Using COc4ccc(c3csc(NC(=O)[C@H]1CCCCN1S(=O)(=O)c2ccc(C)cc2)n3)cc4 at sw.docking.org gave me the following just now: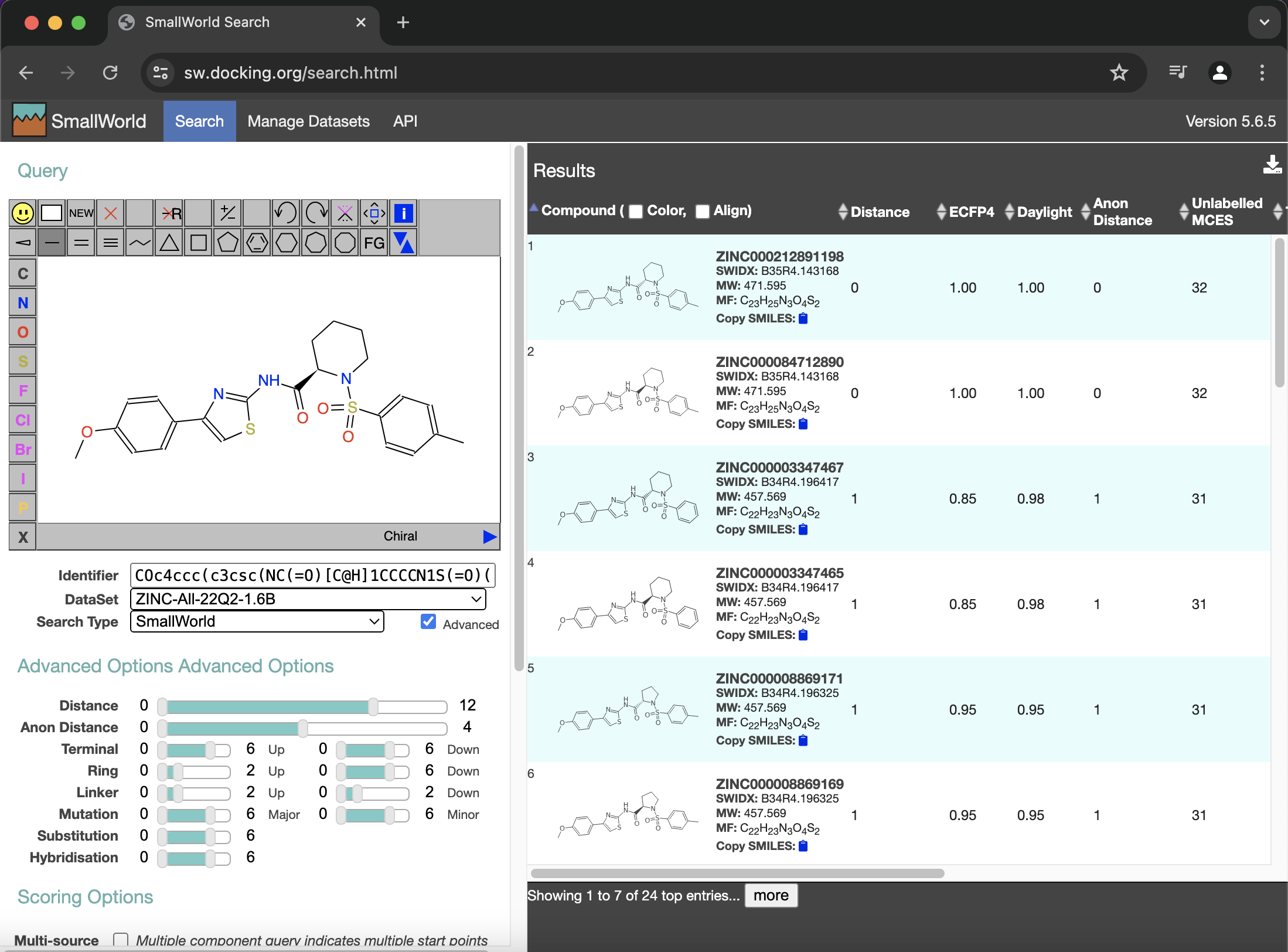
I think hits 1 and 2 are the R and S forms of ML277 (but which is which?). Hits 3 and 4 look like they've removed a methyl (-CH3) from the tolyl group of ML277. I think hits 5 and 6 just changed the twisted 6-membered ring of ML277 to a twisted 5-membered ring. Ideally one should be able to find these top 6 hits on the ZINC site as well, but even a simple search there wasn't working for me earlier today. The Unlabelled MCES column above looks like the # of heavy atoms in each ligand.
Hit 1 above from the sw site gave ZINC ID 212891198. The web page https://zinc20.docking.org/substances/ZINC212891198/ gives more details about it, like Mwt 471.604, logP 4.309, Mol Formula C23H25N3O4S2, 4 rings, 32 heavy atoms, 1 h-bond donor, 6 h-bond acceptors, tPSA 88, and 6 rotatable bonds. If you scroll down, it even gives the ZINC ID 84712890 for its mirror image, which appears at https://zinc20.docking.org/substances/ZINC84712890/. This page for ZINC ID 84712890 even lists KCNQ1, so it is probably the ML277 ligand.
The diagram Sciren posted showing the binding site needs some translation for Foldit.
The numbers posted are 10 residues lower than the PDB model. The "prime" entries like Phe265' mean PDB chain A. Without the prime, like Leu256, it's chain G. So Phe265' refers to PDB chain A, residue 275, as indicated in the segment information window below.
The diagram below shows all the residues identified in the image from Sciren's post, along with the approximate locations of the designations S5', S6', S4-S5, S5, and S6. The ligand shown is just the starting compound, not ML277.
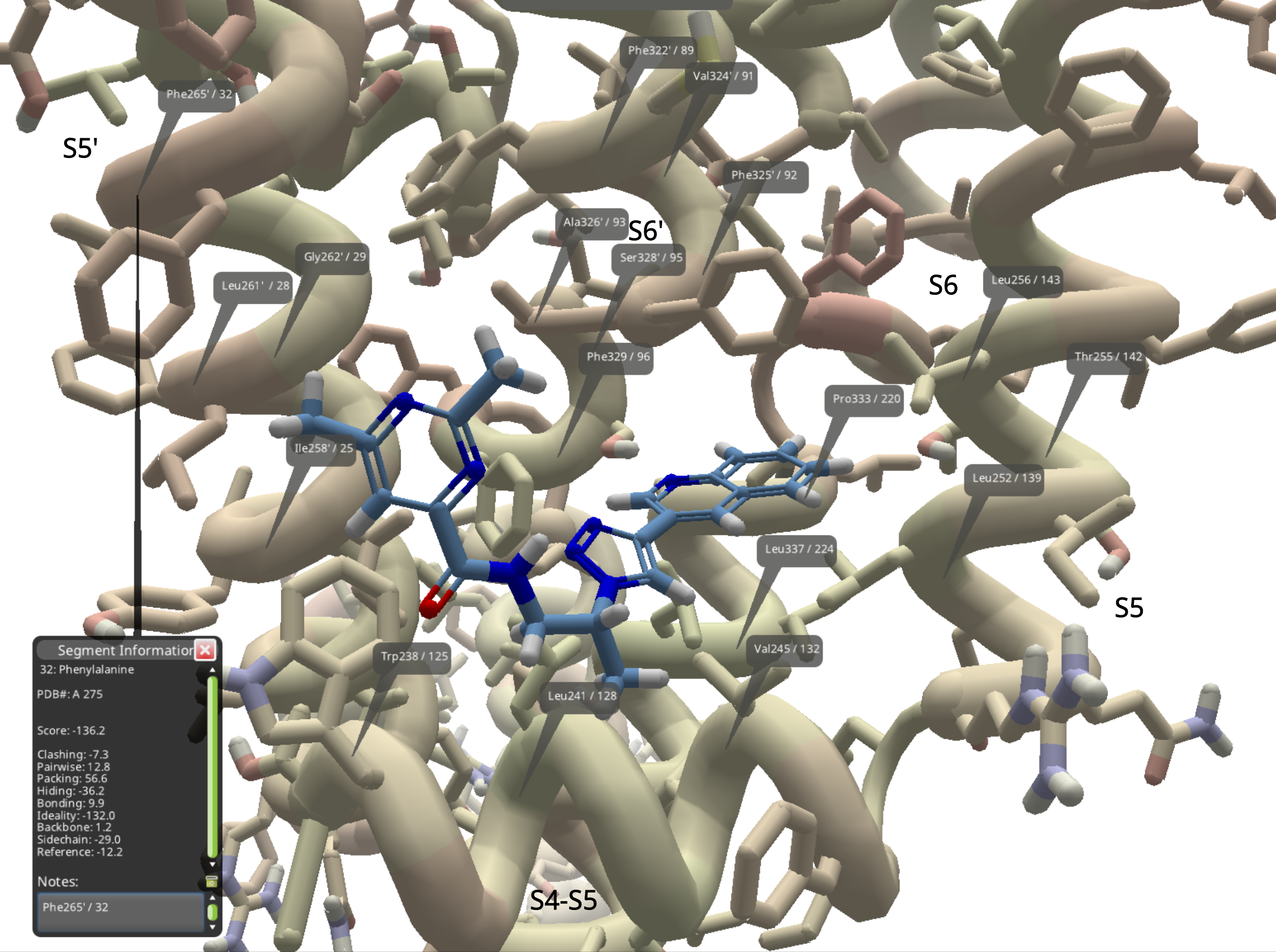
It's not quite as handy as a wiki table, but here are the details, transcribed according to the best of my abilities:
| diagram |
PDB chain |
PDB residue |
Foldit segment |
| Ile258' |
A |
268 |
25 |
| Leu261' |
A |
271 |
28 |
| Gly262' |
A |
272 |
29 |
| Phe265' |
A |
275 |
32 |
| Phe322' |
A |
322 |
89 |
| Val324' |
A |
334 |
91 |
| Phe325' |
A |
335 |
92 |
| Ala326' |
A |
336 |
93 |
| Ser328' |
A |
338 |
95 |
| Phe329' |
A |
339 |
96 |
| Trp238 |
G |
248 |
125 |
| Leu241 |
G |
251 |
128 |
| Val245 |
G |
255 |
132 |
| Leu252 |
G |
262 |
139 |
| Thr255 |
G |
265 |
142 |
| Leu256 |
G |
266 |
143 |
| Pro333 |
G |
343 |
220 |
| Leu337 |
G |
347 |
224 |
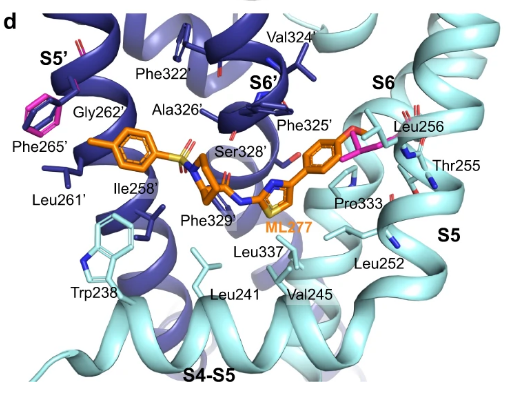
For reference, here's the original diagram from Sciren's post:
.png)
Although the segment information windows show a PDB chain and residue number, nothing identifies the PDB entry. We'll save that for another post, unless someone has already mentioned it.
PDB 6UZZ appears to be a match for the sequence from Foldit. Other PDB matches include 6V00, 6V01, 7XNI, 7XNK, and so on.
In 6UZZ, there are four chains with the same sequence, identified as A, C, E, and G. The Foldit version has a two chains, which are identified with PDB chains A and G in the segment information windows.
6UZZ also has four more chains (B, D, F, and H), which are identified as Calmodulin-1. These chains have a different sequence, and aren't found in the Foldit puzzle. (That Calmodulin part sounds familiar, however.)
I checked the PDB file for 6UZZ, and the residue numbering matches what's shown in the segment information windows in Foldit.
Where Foldit has 241 segment plus the ligand, 6UZZ has a "modelled residue count" of 1,984, and a "deposited residue count" to 2,824. There are lots of missing residues listed in the PDB file. Just be glad to work on the small subset.
I see now that Serca posted 7XNK earlier, with a reference to Calmodulin.
ML277 has a sulfonyl group, which is a little tricky to build in Foldit.
Add sulfonyl group on the wiki shows the step-by-step approach.
I can't wait to make royalties from this amazing project with my team, finally we are treated like we are Protein Folders and not just lab rats :-P
Mmm. Royalties! Much better than rat chow. Can I have some too?
Hi everyone!
Following the discussion, it seems like you all are coming to the correct conclusions. Just to clarify points though:
- ML277 is not the starting compound for the puzzle, but taking its interactions into account could be useful in future testing.
- Throughout the puzzle series we will be using compounds selected from the 900 compounds found in the virtual screening as starting molecules. It will be exciting to learn more about whether your choices match any in those 900 compounds or, more likely, are outside of them. In these endeavors diversity is always highly valued! Either way, the compound library bonus will take effect.
- Thank you @LociOiling! There was indeed a mismatch between the literature diagram and the Foldit representation. In the future I will try and correlate these better.
Thank you all so much, and we are all looking forward to seeing what you can come up with! Speaking of this, just to clarify all compounds created from this puzzle series, like other small molecule puzzle series before it, will enter the public domain so that they can be used for further research purposes. As such there will be no patents or royalties associated with player results.
Sciren




.png)
