
Bruno Kestemont Lv 1
I'v a bug with Ligand queue from reset puzzle, when browsing to fast

Closed since over 2 years ago
Intermediate Intermediate Intermediate Intermediate Intermediate Intermediate Intermediate Intermediate Intermediate Overall Overall Overall Overall Overall Overall Overall Overall Overall Small Molecule Design Small Molecule Design Small Molecule Design Small Molecule Design Small Molecule Design Small Molecule Design Small Molecule Design Small Molecule Design Small Molecule DesignThis puzzle is a followup to the SARS-CoV-2 Nsp3 macrodomain design challenges. Help us rank potential active compounds!
Phase 2 of the CACHE challenge #3 is a ranking of all 1739 molecules which the various participants submitted for testing. In this puzzle you don't have free design with the Small Molecule Design tool, but you instead have the Ligand Queue which will provide you with a small number of possible compounds. You can then use the compound library panel to find related compounds specifically from the library of tested compounds. Then attempt to find the best binding position for the compounds.
Note: To get the most out of the small molecule design tools, we recommend changing you view settings to the Small Molecule Design Preset.
For this round, the ligand queue has a different set of possible compounds (these may get reordered to promote balanced sampling). In addition, we've turned on the Compound Library objective, which should make sure that compounds we're interested in make it to the top of the ranking. - Remember that you may need to submit Ligand Queue compounds to the Compound Library, wait for the results, and then hit wiggle on the compound before the Compound Library objective recognizes the compound. Bounce back and forth between the ligand queue and the compound library search to seek out compounds you think bind best.
Participation in CACHE puzzles is subject to the CACHE Terms of Participation, in particular “the Challenge IP [including Challenge Compounds] will be made freely available in the public domain pursuant to Creative Commons Attribution Only (CC-BY 4.0 or subsequent versions) licensing terms, with the intent that such Challenge IP may be Used and practiced by Users for any purpose”.


I'v a bug with Ligand queue from reset puzzle, when browsing to fast

There is at least one compound in the ligand queue that causes a crash. So far just noted in #bugs-and-feedback on Discord.
I've been able to get the molecular weights, etc., for 28 of the 30 compounds. (I'm not counting the initial compound.)
The comments don't include this information, but the compound library bonus is 4000 points this round.
It sounds like all the starting compounds should be found in the compound library. (Except for the initial compound, with weight 411.414.)
To get the bonus, open the compound library panel and submit the current compound. The results should come back quickly.
Then click "Load Library". The library contents should appear in a few moments. The first result should be "exact match".
If you see "exact match", all you need to do shake for a second, followed by a brief wiggle. You should now have the compound library bonus.
Another option if you have the compound saved is to reload it from Open/Share Solutions when you see the "exact match". This should also get the bonus.
You do not need to click "Accept Compound" to get the bonus.
In general, if you have the compound library open when you load a solution, you should get the bonus immediately if it's in the results for a compound you submitted.
If you have the compound library open, you can start the ligand queue, and then submit library searches as you go. The compound library must be opened before the ligand queue. Unfortunately, you can't save a solution, shake, or wiggle while the ligand queue is open.
You can get the molecular weight and other properties in the "Small Molecule Properties" window, which opens with the "v" hotkey when the ligand is selected. You'll need to close or more the "Small Molecule View Options" window to see "Small Molecule Properties".

Official bug report on the ligand queue crash: https://fold.it/forum/bugs/ligand-queue-crash-is-back-for-2363

I'm getting compounds from ligand queue without library bonus. Current example has 2 library entries having similarity of 1.0 with the molecule from ligand queue. Maybe a mismatch in conformer generation -> regenerating smiles -> similarity calculation?
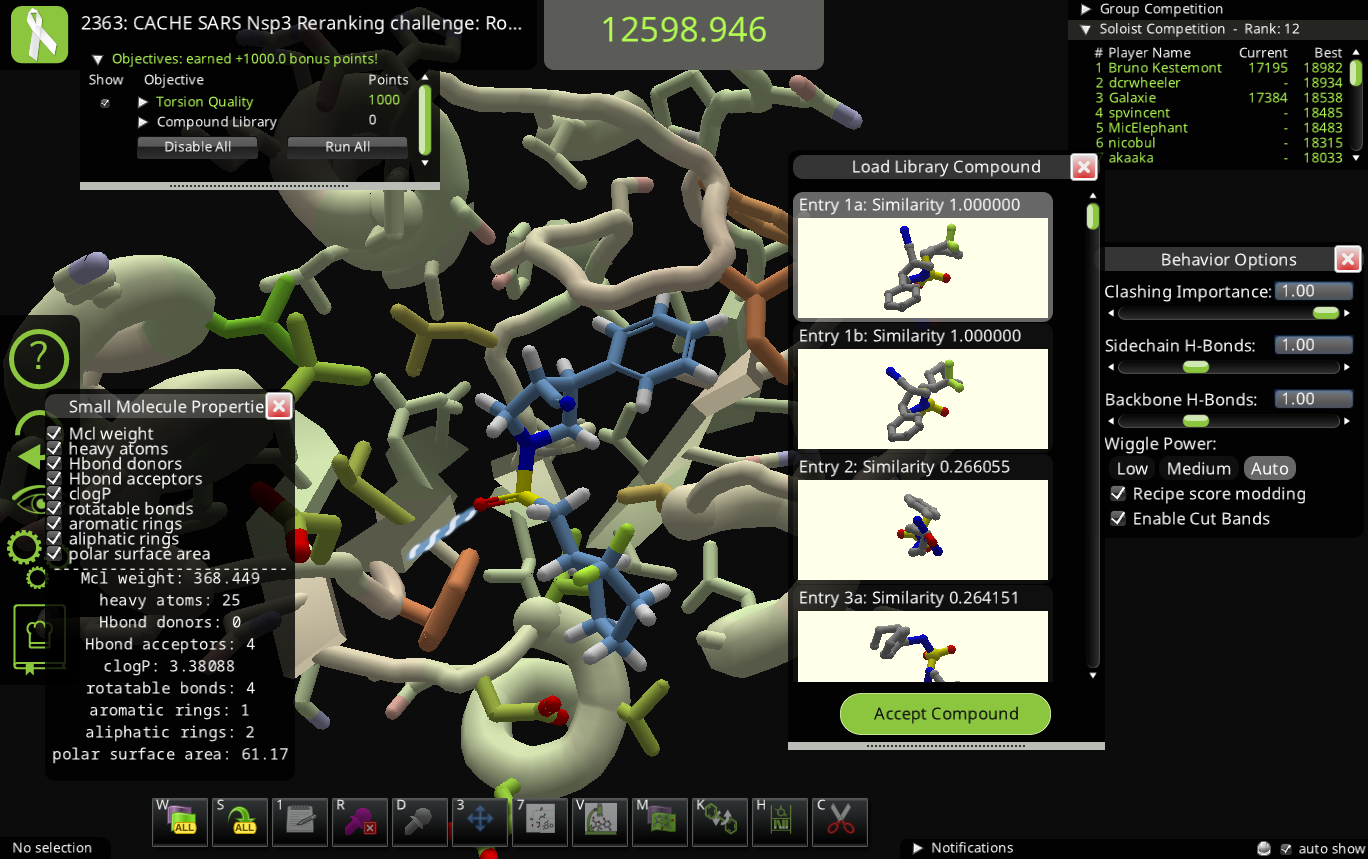
(edit: add screenshot)

Another way to get the Library bonus: save locally. Open library, accept one of them. Load from local save.

I added a bug report on the no-bonus issue: https://fold.it/forum/bugs/puzzle-2363-compounds-385-445-and-385-552-don-t-get-cl-bonus

Update: as @jeff101 noted in #veteran today, we now have two additional compounds, giving a total of 30 plus the "initial" compound. The crashing problem has been resolved, but you should probably restart Foldit or at least reload the puzzle to make sure you have the current puzzle setup.
Here are the properties for the 30 compounds in the ligand queue. The "initial" compound with weight 411.414 it omitted, based on the comments for puzzle 2360. I also left out the aromatic and aliphatic ring counts on account of space.
| # | weight | cLogP | PSA | heavy atoms | H-bond donors | H-bond acceptors | rotatable bonds |
|---|---|---|---|---|---|---|---|
| 1 | 204.042 | 2.62990 | 12.03 | 10 | 1 | 1 | 1 |
| 2 | 218.300 | 1.52980 | 36.36 | 16 | 1 | 3 | 1 |
| 3 | 231.215 | 0.23572 | 78.22 | 17 | 0 | 7 | 2 |
| 4 | 246.314 | 1.29920 | 58.95 | 18 | 2 | 5 | 5 |
| 5 | 262.357 | 2.59240 | 51.45 | 19 | 1 | 5 | 7 |
| 6 | 264.329 | 1.00380 | 76.02 | 19 | 1 | 6 | 5 |
| 7 | 271.263 | 2.19384 | 59.16 | 19 | 1 | 4 | 4 |
| 8 | 281.337 | 0.24970 | 78.71 | 19 | 2 | 6 | 3 |
| 9 | 290.319 | 2.14098 | 64.33 | 20 | 1 | 4 | 3 |
| 10 | 291.250 | 2.76390 | 60.17 | 19 | 0 | 4 | 3 |
| 11 | 297.366 | 1.65440 | 86.38 | 22 | 2 | 7 | 7 |
| 12 | 298.318 | 0.66080 | 98.58 | 21 | 3 | 7 | 6 |
| 13 | 299.274 | 0.59650 | 92.18 | 19 | 2 | 6 | 7 |
| 14 | 306.395 | 0.65840 | 86.80 | 21 | 2 | 7 | 6 |
| 15 | 313.327 | 3.46480 | 79.62 | 22 | 4 | 5 | 4 |
| 16 | 328.368 | 1.76750 | 71.89 | 24 | 1 | 6 | 7 |
| 17 | 343.261 | 1.16010 | 105.17 | 24 | 2 | 7 | 7 |
| 18 | 343.361 | -0.36368 | 127.45 | 23 | 2 | 9 | 10 |
| 19 | 368.449 | 3.38088 | 61.17 | 25 | 0 | 4 | 4 |
| 20 | 376.428 | 1.51610 | 115.40 | 28 | 2 | 10 | 2 |
| 21 | 380.426 | -0.26300 | 93.85 | 26 | 0 | 9 | 3 |
| 22 | 385.445 | 3.36860 | 79.81 | 27 | 0 | 7 | 7 |
| 23 | 385.552 | 3.88500 | 54.34 | 28 | 1 | 5 | 8 |
| 24 | 388.424 | 2.57140 | 96.26 | 27 | 2 | 7 | 8 |
| 25 | 390.417 | 1.32140 | 118.32 | 27 | 2 | 8 | 8 |
| 26 | 396.245 | 3.06470 | 66.39 | 23 | 0 | 3 | 3 |
| 27 | 410.358 | 2.69010 | 40.62 | 28 | 0 | 4 | 6 |
| 28 | 421.457 | 1.55060 | 126.81 | 31 | 2 | 9 | 5 |
| 29 | 433.512 | 2.96514 | 97.22 | 32 | 3 | 8 | 7 |
| 30 | 451.918 | 3.56834 | 102.31 | 32 | 0 | 9 | 8 |

Here are the SMILES strings for each compound, now updated for all 30 compounds. The SMILES came from hand transcriptions of the string shown for each of the ligand queue entries. Errors are possible, but generally transcription errors mean an invalid SMILES string. All of these strings at least loaded into Jmol and produced the chemical formulas shown here.
| # | weight | SMILES | formula |
|---|---|---|---|
| 1 | 204.042 | CNc1c(F)cccc1Br |
C7H7BrFN |
| 2 | 218.300 | CC1C(O)CN1c1ccnc2c1CCCC2 |
C13H18N2O |
| 3 | 231.215 | Cc1cc(Cn2nc3cnccn3c2=O)on1 |
C10H11N5O2 |
| 4 | 246.314 | C#CCNC(=O)NC1CCCCC1n1cccn1 |
C13H18N4O |
| 5 | 262.357 | CCC(CC)C(CNc1cccc2nncn12)OC |
C14H22N4O |
| 6 | 264.329 | CC(C)Cn1cc(NC(=O)C2CCNC(=O)C2)cn1 |
C13H20N4O2 |
| 7 | 271.263 | Cc1cc(C(=O)OCC2CC(F)(F)C2)c(=O)[nH]c1C |
C13H15F2NO3 |
| 8 | 281.337 | OC[C@@H]1CN(c2nc3cccnc3s2)C[C@@H](CO)O1 |
C12H15N3O3S |
| 9 | 290.319 | N#CCC1(O)CN(C(=O)c2cc3cc(F)ccc3s2)C1 |
C14H11FN2O2S |
| 10 | 291.250 | CS(=O)(=O)c1cccc(-c2nc(C(F)(F)F)co2)c1 |
C11H8F3NO3S |
| 11 | 297.366 | CN(CCCc1cc(-c2ccccc2)n[nH]1)Cc1nn[nH]n1 |
C15H19N7 |
| 12 | 298.318 | CC(O)CNc1cc(N2C[C@@H](F)C[C@H]2CC(=O)O)ncn1 |
C13H19FN4O3 |
| 13 | 299.274 | O=S(=O)(CCC(F)(F)F)NCCc1cc(O)ncn1 |
C9H12F3N3O3S |
| 14 | 306.395 | CN(CC(=O)NCC1(C)CSC1)c1ncnc2nc[nH]c12 |
C13H18N6OS |
| 15 | 313.327 | Nc1nc(NCC2(C(F)(F)F)CCCCC2)c2cc[nH]c2n1 |
C14H18F3N5 |
| 16 | 328.368 | O=C(COCCOc1ccccc1)N1CCc2ccnc(O)c2C1 |
C18H20N2O4 |
| 17 | 343.261 | O=C(OCCc1cnc(=O)[nH]c1)c1cc(C(O)C(F)(F)F)ccn1 |
C14H12F3N3O4 |
| 18 | 343.361 | COC(=O)CCNC(=O)CSc1nc(=O)[nH]c(C)c1C(=O)OC |
C13H17N3O6S |
| 19 | 368.449 | N#C[C@@H]1CN(S(=O)CC2CCCCC2(F)F)C[C@H]1c1ccccc1 |
C18H22F2N2OS |
| 20 | 376.428 | c1nc(N2CCC3(CC2)CCN(c2ncnc4[nH]ncc24)C3)c2cn[nH]c2n1 |
C18H20N10 |
| 21 | 380.426 | COc1ccc2c(c1)N(C)CCN2S(=O)(=O)c1cn(C)c(=O)n(C)c1=O |
C16H20N4O5S |
| 22 | 385.445 | CCN(C(C)=O)c1nc(C=CC(=O)c2cccc(N3CCOC3=O)c2)cs1 |
C19H19N3O4S |
| 23 | 385.552 | CCCCC1CCC(C(=O)N2CCC(NC(=O)c3ccn(C)c3)C23CC3)CC1 |
C23H35N3O2 |
| 24 | 388.424 | COc1ccc(OCc2nnc(SCC(N)=O)n2-c2ccccc2F)cc1 |
C18H17FN4O3S |
| 25 | 390.417 | O=C(C=Cc1ccc[nH]1)O[C@H]1CS(=O)(=O)C[C@@H]1OC(=O)C=Cc1ccc[nH]1 |
C18H18N2O6S |
| 26 | 396.245 | O=C(c1nc(Br)c2c(F)cccn12)N1CCCC1c1nncs1 |
C14H11BrFN5OS |
| 27 | 410.358 | O=C(N1C[C@H]2[C@@H](C1)[C@@H]2C(F)(F)F)C1(C(=O)N2C[C@H]3[C@@H](C2)[C@@H]3C(F)(F)F)CCC1 |
C18H20F6N2O2 |
| 28 | 421.457 | CCn1c(=O)[nH]c(=O)c2c(C(=O)NC3CCc4cnccc4C3=O)cc(C(C)C)nc21 |
C22H23N5O4 |
| 29 | 433.512 | Cc1cc2ncn(CC(O)CNCc3c[nH]nc3-c3ccc4c(c3)OCCO4)c2cc1C |
C24H27N5O3 |
| 30 | 451.918 | Cc1nc2ncnn2c(C)c1CCC(=O)N(Cc1nnc(-c2ccc(Cl)cc2)o1)C1CC1 |
C22H22ClN7O2 |
Sorry about the formatting, I haven't found a way to control table properties in Foldit's implementation of Markdown.
I did find a way to escape the @ signs in SMILES string to avoid creating spurious links. Add a backtick around the SMILES string did the trick. There must be a space between the backtick and the vertical bar used to delimit the table columns.

The issue with the ligand in the ligand queue which is causing a crash should be fixed. You may need to restart your client to pull in the updated puzzle files.

Looks like the issue with the ligand queue crash was also affecting the compound library search. I've adjusted a few things, and I think things are a bit better now.
You may (will) have to resubmit compounds to the library to take advantage of the fixed compound library search.